Comparing Alignment Methods
Source:vignettes/articles/alignment-comparison.Rmd
alignment-comparison.RmdIntroduction
fdars provides four approaches to curve alignment, each with different trade-offs in terms of automation, smoothness, interpretability, and speed. This article compares them side-by-side on the same data, and provides guidance on when to use each method.
The four methods are:
-
Elastic alignment (
elastic.align,karcher.mean) – global optimization via dynamic programming in SRSF space. Seevignette("elastic-alignment", package = "fdars")for full details. -
Landmark registration
(
landmark.register) – feature-based piecewise-linear warping. Seevignette("landmark-registration", package = "fdars")for full details. -
Constrained elastic alignment
(
elastic.align.constrained) – elastic alignment with landmark pass-through constraints -
TSRVF (
tsrvf.transform) – not an alignment method per se, but a linearization of elastic alignment for downstream analysis. Seevignette("tsrvf", package = "fdars")for full details.
Test Data
We use two datasets that highlight different alignment challenges.
Dataset 1: Smooth Phase Shifts
Curves with smooth, global timing differences – the classic alignment scenario:
set.seed(42)
argvals <- seq(0, 1, length.out = 200)
n <- 15
data_smooth <- matrix(0, n, 200)
for (i in 1:n) {
shift <- runif(1, -0.1, 0.1)
amp <- rnorm(1, 1, 0.15)
data_smooth[i, ] <- amp * sin(2 * pi * (argvals - shift)) +
0.4 * amp * sin(4 * pi * (argvals - shift * 0.7))
}
fd_smooth <- fdata(data_smooth, argvals = argvals)Dataset 2: Distinct Feature Shifts
Curves with clearly identifiable peaks that shift independently – where domain knowledge matters:
data_feat <- matrix(0, n, 200)
for (i in 1:n) {
pk1 <- runif(1, 0.15, 0.35)
pk2 <- runif(1, 0.55, 0.75)
w1 <- rnorm(1, 1, 0.2)
w2 <- rnorm(1, 0.8, 0.15)
data_feat[i, ] <- w1 * exp(-200 * (argvals - pk1)^2) +
w2 * exp(-200 * (argvals - pk2)^2) +
0.05 * rnorm(200)
}
fd_feat <- fdata(data_feat, argvals = argvals)
p1 <- plot(fd_smooth) + labs(title = "Dataset 1: Smooth Phase Shifts")
p2 <- plot(fd_feat) + labs(title = "Dataset 2: Distinct Features")
p1 + p2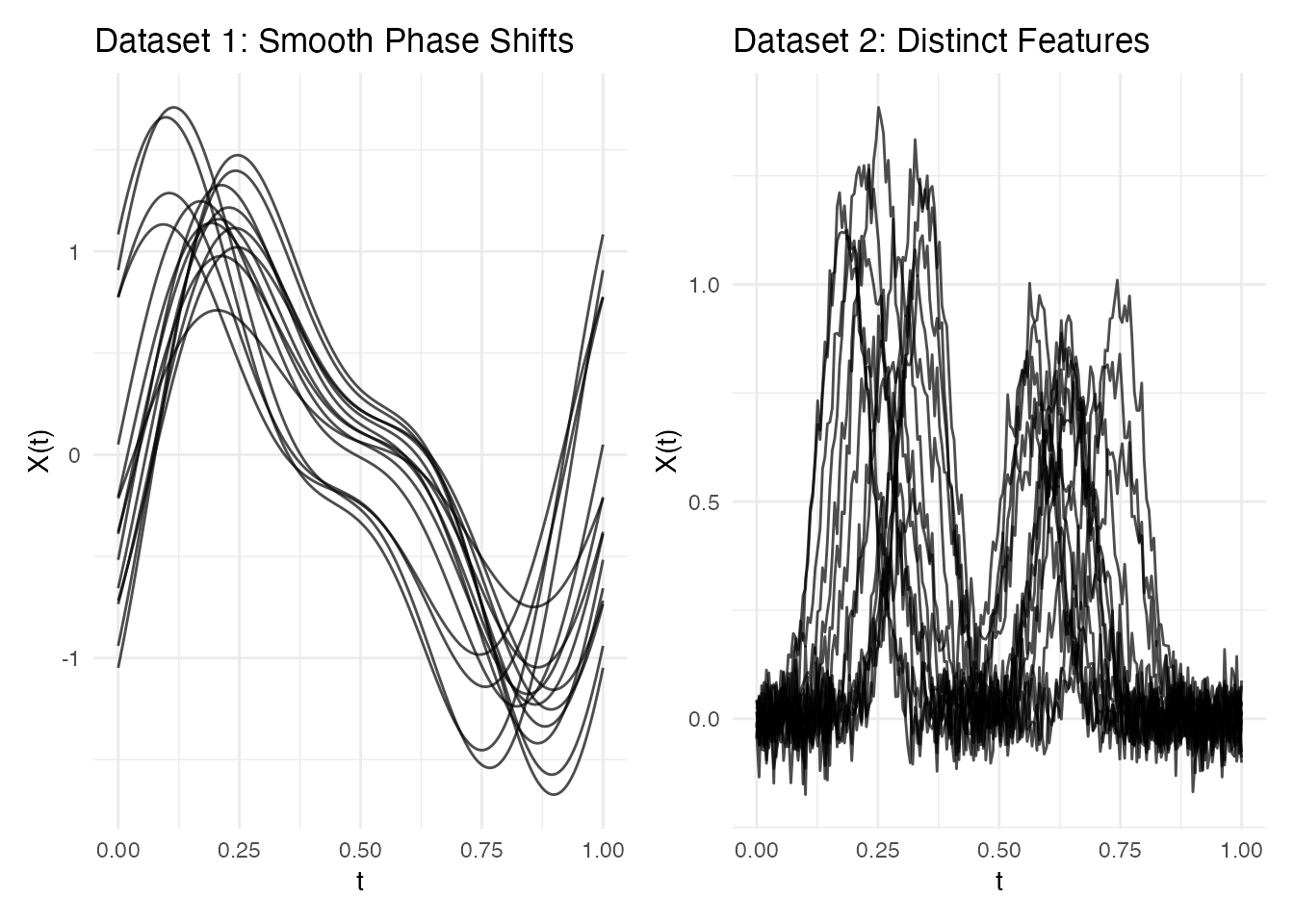
Method 1: Elastic Alignment
Elastic alignment finds the globally optimal warping function by minimizing the distance between SRSFs via dynamic programming. It requires no feature detection and produces smooth, diffeomorphic warping functions.
Mathematical basis: minimize over the warping group , where is the SRSF of .
ea_smooth <- elastic.align(fd_smooth)
ea_feat <- elastic.align(fd_feat)
p1 <- plot(ea_smooth, type = "aligned") + labs(title = "Elastic (smooth data)")
p2 <- plot(ea_feat, type = "aligned") + labs(title = "Elastic (feature data)")
p1 + p2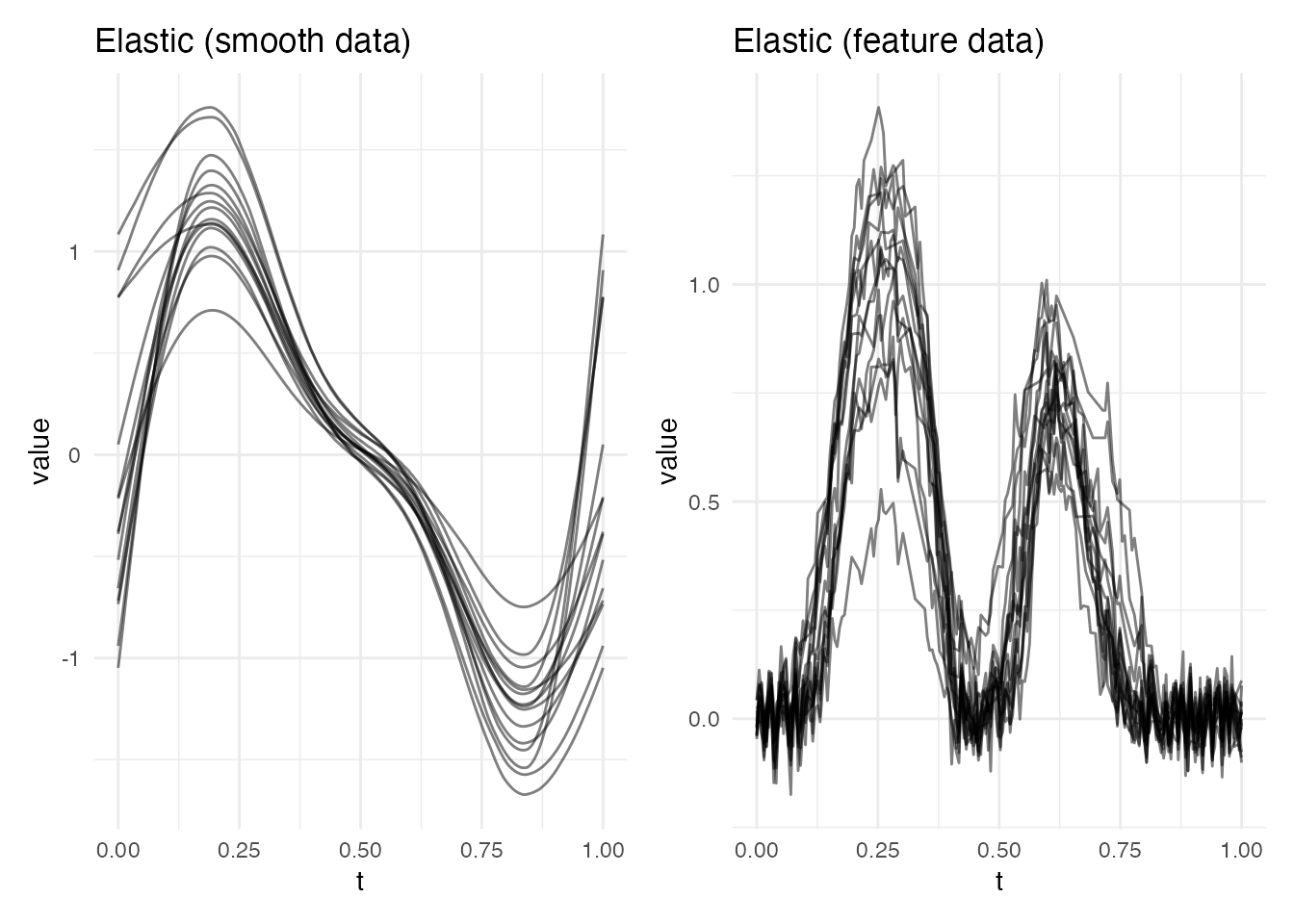
Method 2: Landmark Registration
Landmark registration detects features (peaks, valleys, etc.) and warps curves so that corresponding features align to common target positions. The warping is piecewise-linear between anchor points.
Mathematical basis: For landmarks and targets , construct as a piecewise-linear function satisfying .
lr_smooth <- landmark.register(fd_smooth, kind = "peak", min.prominence = 0.3,
expected.count = 1)
lr_feat <- landmark.register(fd_feat, kind = "peak", min.prominence = 0.2,
expected.count = 2)
p1 <- plot(lr_smooth, type = "registered") +
labs(title = "Landmark (smooth data)")
p2 <- plot(lr_feat, type = "registered") +
labs(title = "Landmark (feature data)")
p1 + p2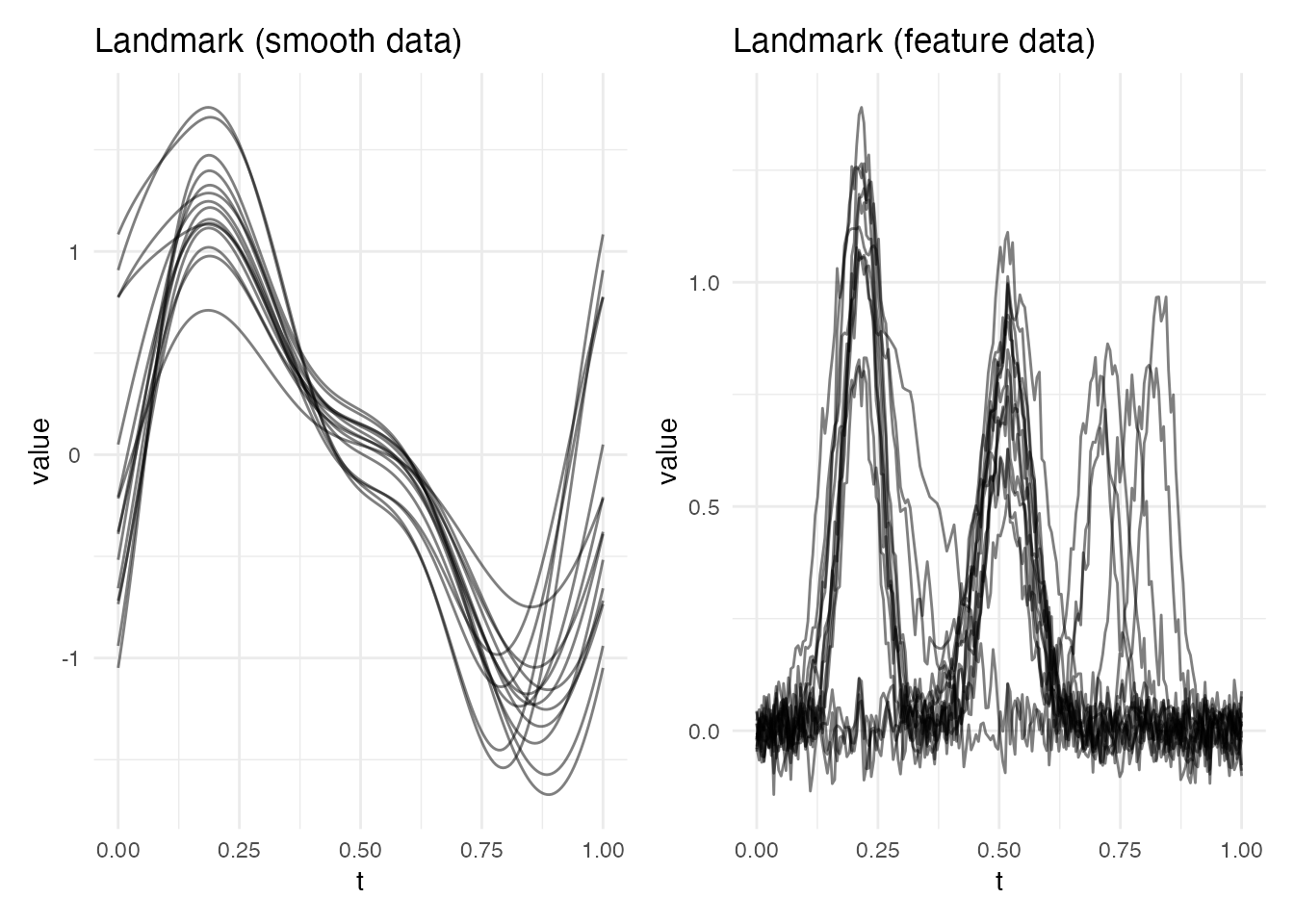
Method 3: Constrained Elastic Alignment
Constrained elastic alignment combines the smooth, globally optimal warping of elastic alignment with landmark pass-through constraints. The warping is computed by segmented dynamic programming between landmark anchor points.
Mathematical basis: minimize subject to for each landmark pair.
ec_smooth <- elastic.align.constrained(fd_smooth, kind = "peak",
min.prominence = 0.3,
expected.count = 1)
ec_feat <- elastic.align.constrained(fd_feat, kind = "peak",
min.prominence = 0.2,
expected.count = 2)
p1 <- plot(ec_smooth, type = "aligned") +
labs(title = "Constrained (smooth data)")
p2 <- plot(ec_feat, type = "aligned") +
labs(title = "Constrained (feature data)")
p1 + p2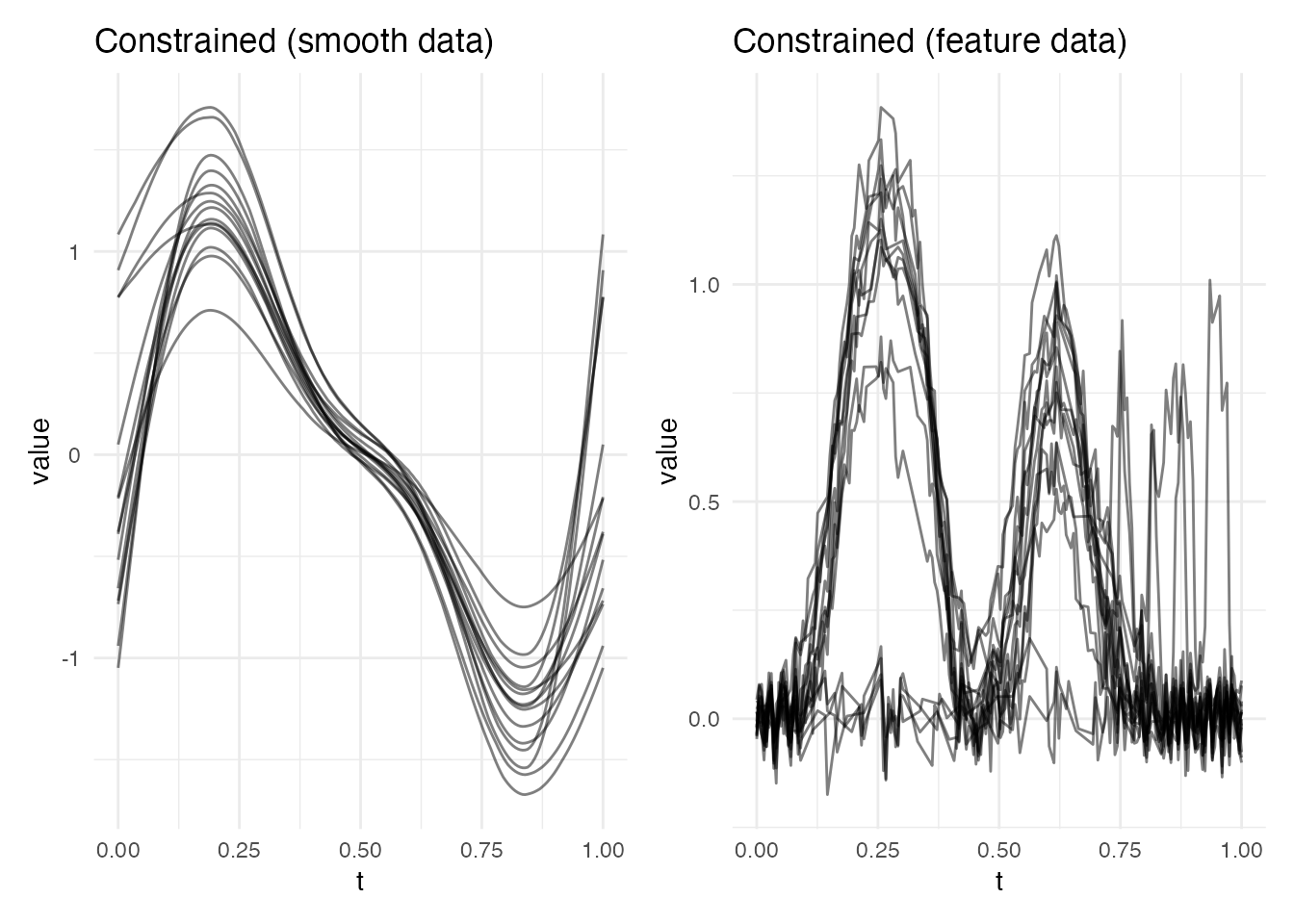
Warping Function Comparison
The warping functions reveal the fundamental difference between methods. Elastic produces smooth warps, landmark produces piecewise-linear warps, and constrained is smooth but passes through landmark anchor points:
p1 <- plot(ea_feat, type = "warps") + labs(title = "Elastic Warps")
p2 <- plot(lr_feat, type = "warps") + labs(title = "Landmark Warps")
p3 <- plot(ec_feat, type = "warps") + labs(title = "Constrained Warps")
p1 + p2 + p3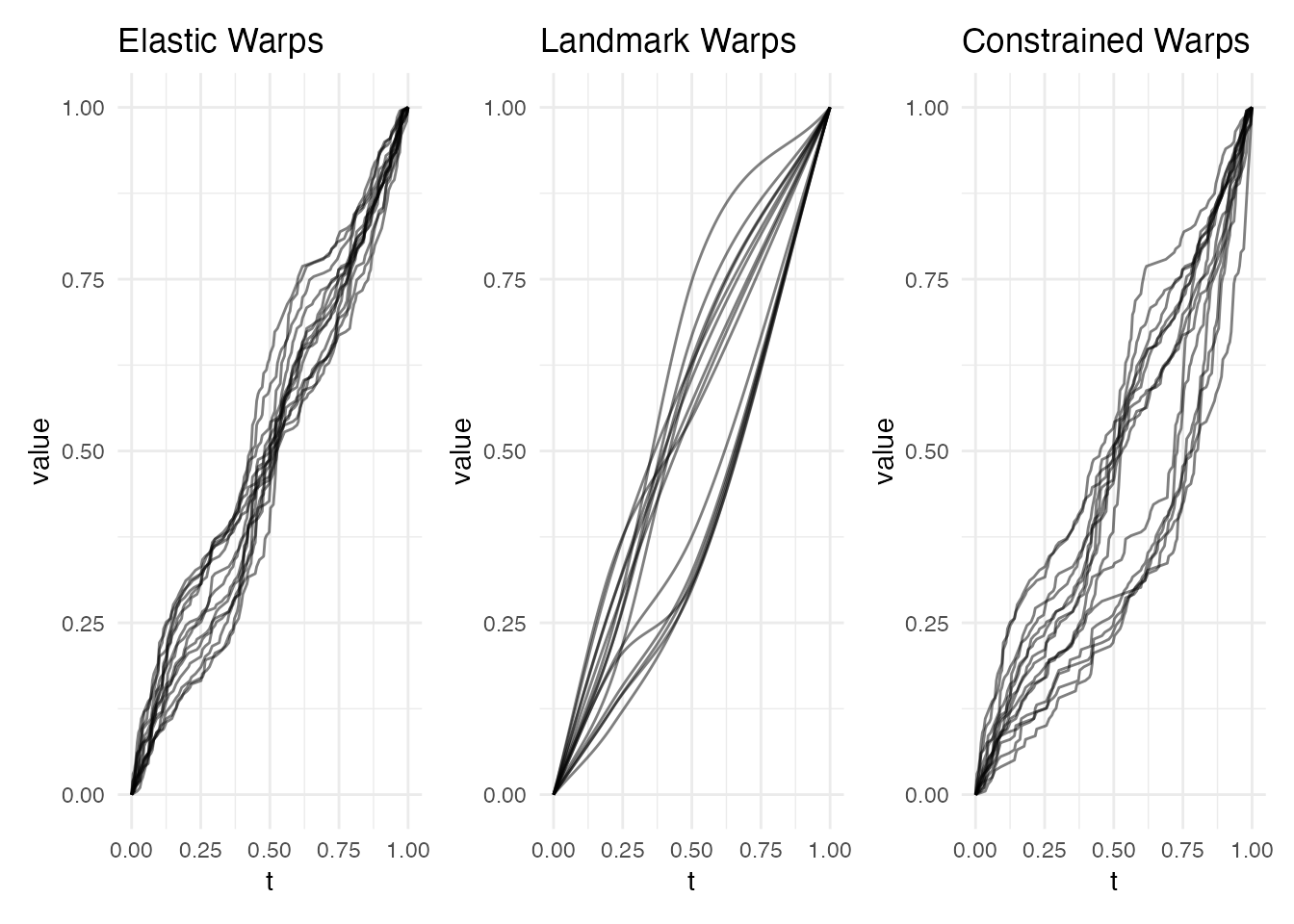
Quantitative Comparison
Variance Reduction
Variance reduction (VR) measures how much pointwise variance is removed by alignment. Higher VR means more phase variability was captured:
vr <- function(original, aligned) {
var_orig <- mean(apply(original$data, 2, var))
var_aligned <- mean(apply(aligned$data, 2, var))
1 - var_aligned / var_orig
}
results <- data.frame(
Dataset = rep(c("Smooth shifts", "Distinct features"), each = 3),
Method = rep(c("Elastic", "Landmark", "Constrained"), 2),
VR = c(
vr(fd_smooth, ea_smooth$aligned),
vr(fd_smooth, lr_smooth$registered),
vr(fd_smooth, ec_smooth$aligned),
vr(fd_feat, ea_feat$aligned),
vr(fd_feat, lr_feat$registered),
vr(fd_feat, ec_feat$aligned)
)
)
results$VR <- round(results$VR, 3)
print(results)
#> Dataset Method VR
#> 1 Smooth shifts Elastic 0.527
#> 2 Smooth shifts Landmark 0.380
#> 3 Smooth shifts Constrained 0.527
#> 4 Distinct features Elastic 0.718
#> 5 Distinct features Landmark 0.248
#> 6 Distinct features Constrained 0.054
ggplot(results, aes(x = Method, y = VR, fill = Dataset)) +
geom_col(position = "dodge") +
labs(title = "Variance Reduction by Method",
y = "Variance Reduction (higher = better)") +
scale_fill_manual(values = c("Smooth shifts" = "steelblue",
"Distinct features" = "coral")) +
ylim(0, 1)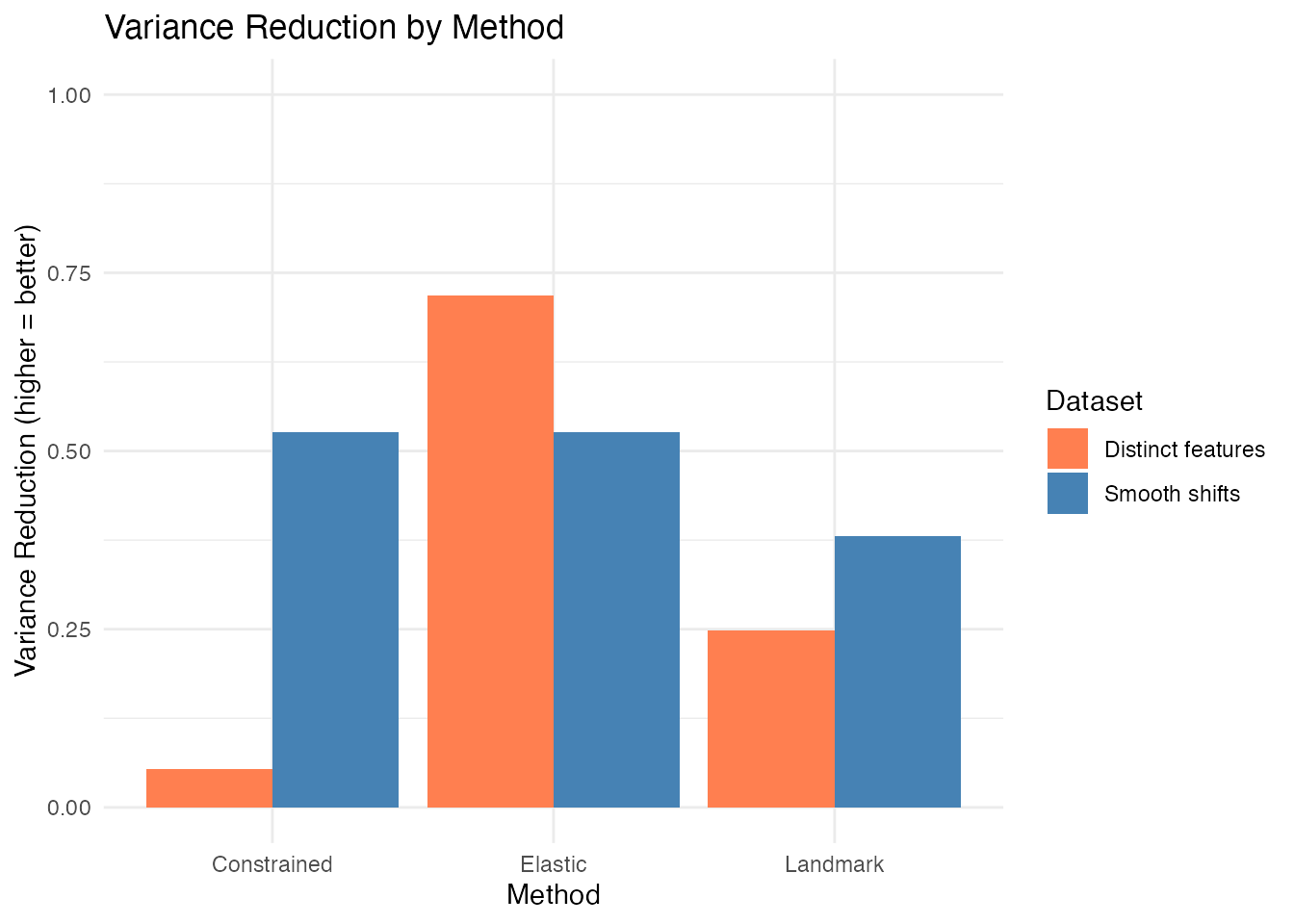
Alignment Quality Diagnostics
For the Karcher mean (elastic), we can compute detailed diagnostics:
km_smooth <- karcher.mean(fd_smooth, max.iter = 15)
aq_smooth <- alignment.quality(fd_smooth, km_smooth)
print(aq_smooth)
#> Alignment Quality Diagnostics
#> Mean warp complexity: 0.1567
#> Mean warp smoothness: 127.4957
#> Total variance: 0.1426
#> Amplitude variance: 0.067
#> Phase variance: 0.0757
#> Phase/Total ratio: 0.5306
#> Mean VR: 0.4609Amplitude-Phase Decomposition
Elastic alignment uniquely provides a principled decomposition of total variability into amplitude and phase components:
plot(aq_smooth, type = "variance")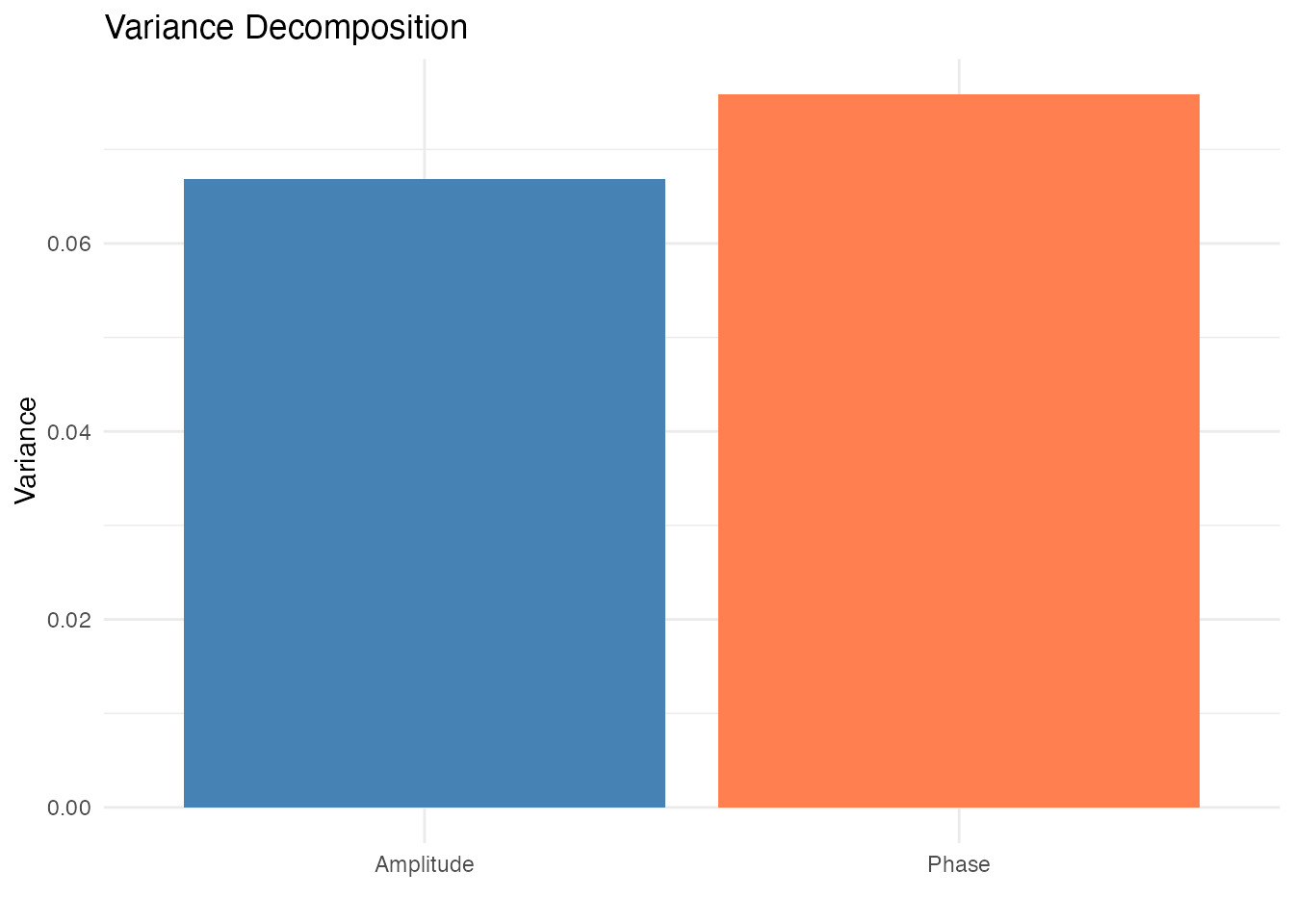
When Methods Succeed and Fail
The examples above use well-behaved data. Real data is messier. This section shows targeted scenarios where each method shines and where it breaks down, so you can match the method to your data.
Scenario 1: Elastic Excels — Smooth Global Warping
When curves differ only by smooth, continuous timing changes with no distinct features, elastic alignment is ideal. It finds the optimal warping without needing any manual tuning:
set.seed(101)
argvals <- seq(0, 1, length.out = 200)
n <- 12
# Smooth curves with continuous phase variation
data_global <- matrix(0, n, 200)
for (i in 1:n) {
# Smooth nonlinear warp of the time axis
warp_strength <- runif(1, -0.15, 0.15)
t_warped <- argvals + warp_strength * sin(pi * argvals)
t_warped <- (t_warped - min(t_warped)) / (max(t_warped) - min(t_warped))
amp <- rnorm(1, 1, 0.1)
data_global[i, ] <- amp * (sin(2 * pi * t_warped) + 0.3 * cos(6 * pi * t_warped))
}
fd_global <- fdata(data_global, argvals = argvals)
ea_global <- elastic.align(fd_global)
# Landmark registration struggles: no clear isolated peaks
lr_global <- tryCatch(
landmark.register(fd_global, kind = "peak", min.prominence = 0.3, expected.count = 1),
error = function(e) NULL
)
p1 <- plot(fd_global) + labs(title = "Original: Smooth Warping")
p2 <- plot(ea_global, type = "aligned") + labs(title = "Elastic: Clean alignment")
if (!is.null(lr_global)) {
p3 <- plot(lr_global, type = "registered") + labs(title = "Landmark: Misses subtlety")
p1 + p2 + p3 + plot_layout(ncol = 3)
} else {
p3 <- ggplot() + annotate("text", x = 0.5, y = 0.5,
label = "Landmark failed:\nno consistent features found",
size = 4, hjust = 0.5) + theme_void() + labs(title = "Landmark: Failed")
p1 + p2 + p3 + plot_layout(ncol = 3)
}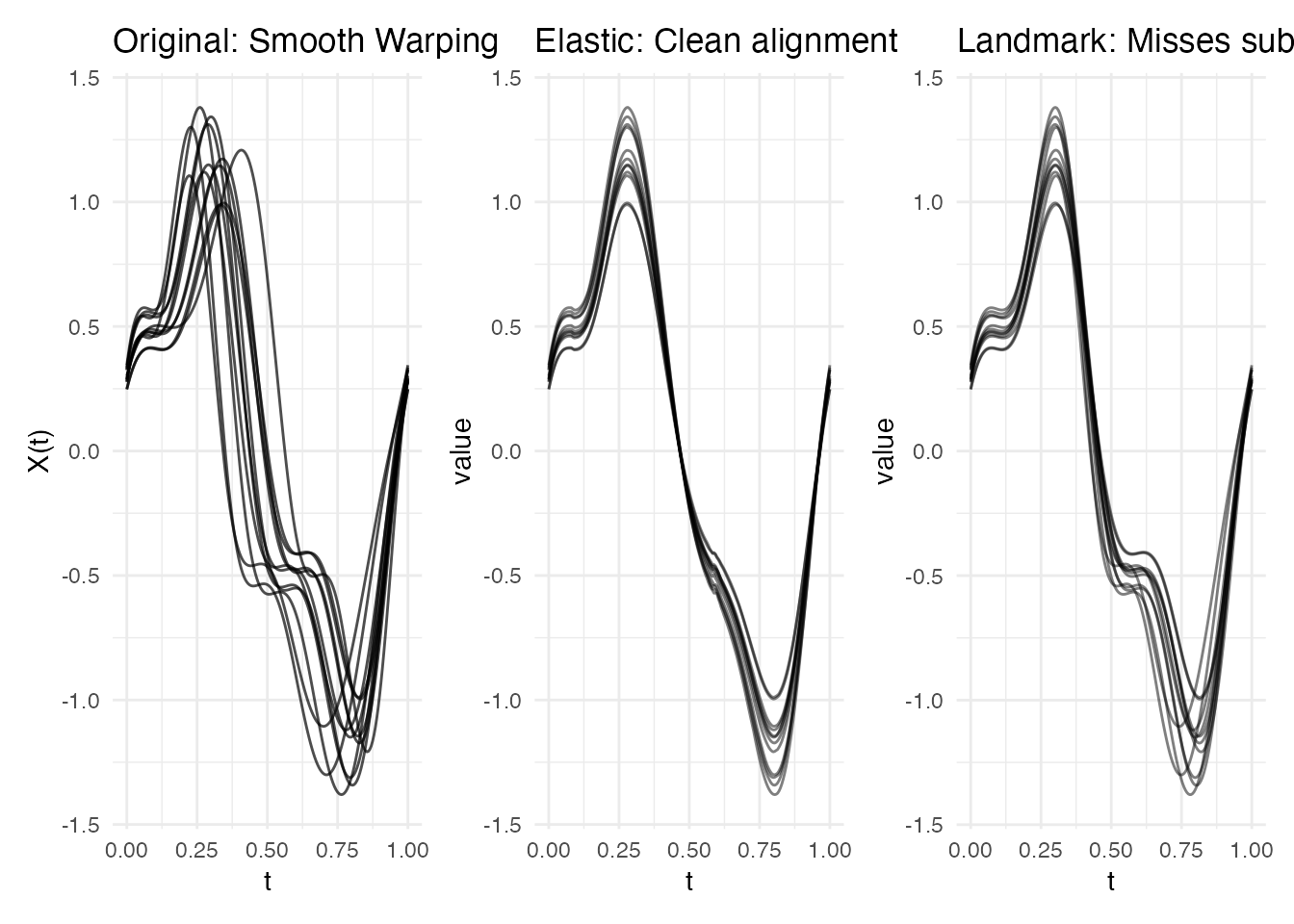
Why elastic wins: The Fisher-Rao metric captures continuous phase variation naturally. Landmark registration needs identifiable features, and these smooth curves have overlapping, broad oscillations with no clear anchor points.
Scenario 2: Landmark Excels — Identifiable Features Shifting Independently
When curves have clearly defined peaks or valleys that shift to different positions, landmark registration directly targets what matters:
# Two-peak curves where peaks shift independently
data_peaks <- matrix(0, n, 200)
for (i in 1:n) {
pk1_pos <- runif(1, 0.15, 0.35)
pk2_pos <- runif(1, 0.6, 0.8)
h1 <- rnorm(1, 1.5, 0.2)
h2 <- rnorm(1, 1.0, 0.15)
data_peaks[i, ] <- h1 * exp(-300 * (argvals - pk1_pos)^2) +
h2 * exp(-300 * (argvals - pk2_pos)^2)
}
fd_peaks <- fdata(data_peaks, argvals = argvals)
lr_peaks <- landmark.register(fd_peaks, kind = "peak", min.prominence = 0.3,
expected.count = 2)
ea_peaks <- elastic.align(fd_peaks)
p1 <- plot(fd_peaks) + labs(title = "Original: Two Shifting Peaks")
p2 <- plot(lr_peaks, type = "registered") + labs(title = "Landmark: Peaks aligned")
p3 <- plot(ea_peaks, type = "aligned") + labs(title = "Elastic: May swap features")
p1 + p2 + p3 + plot_layout(ncol = 3)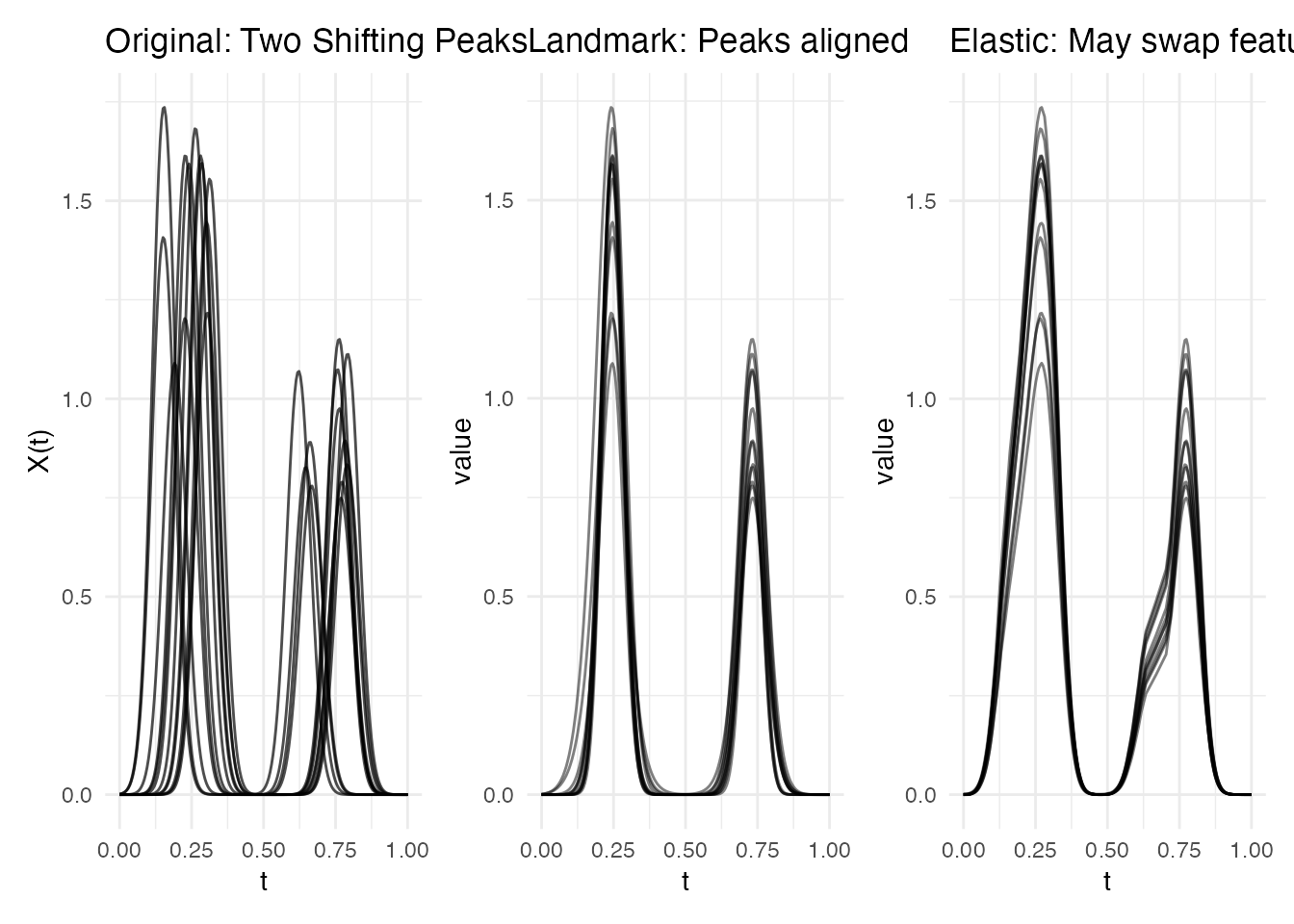
Why landmark wins: It guarantees that peak 1 aligns to peak 1 and peak 2 to peak 2. Elastic alignment has no feature awareness — it minimizes overall distance, which can sometimes swap or merge features when they are far apart.
Scenario 3: Elastic Struggles — Noisy Data
When curves are noisy, elastic alignment’s dynamic programming can over-warp, matching noise spikes rather than underlying signal. The solution: smooth first, then align:
# Clean signal + heavy noise
data_noisy <- matrix(0, n, 200)
for (i in 1:n) {
shift <- runif(1, -0.08, 0.08)
signal <- sin(2 * pi * (argvals - shift))
data_noisy[i, ] <- signal + rnorm(200, 0, 0.4)
}
fd_noisy <- fdata(data_noisy, argvals = argvals)
# Direct alignment on noisy data: over-warping
ea_noisy_raw <- elastic.align(fd_noisy)
# Better approach: smooth first, then align
fd_smooth_noisy <- fdata(t(apply(fd_noisy$data, 1, function(y) {
stats::smooth.spline(argvals, y, spar = 0.6)$y
})), argvals = argvals)
ea_noisy_smooth <- elastic.align(fd_smooth_noisy)
p1 <- plot(fd_noisy) + labs(title = "Original: Noisy Curves")
p2 <- plot(ea_noisy_raw, type = "aligned") +
labs(title = "Elastic on raw: Over-warps")
p3 <- plot(ea_noisy_smooth, type = "aligned") +
labs(title = "Smooth then align: Controlled")
p1 + p2 + p3 + plot_layout(ncol = 3)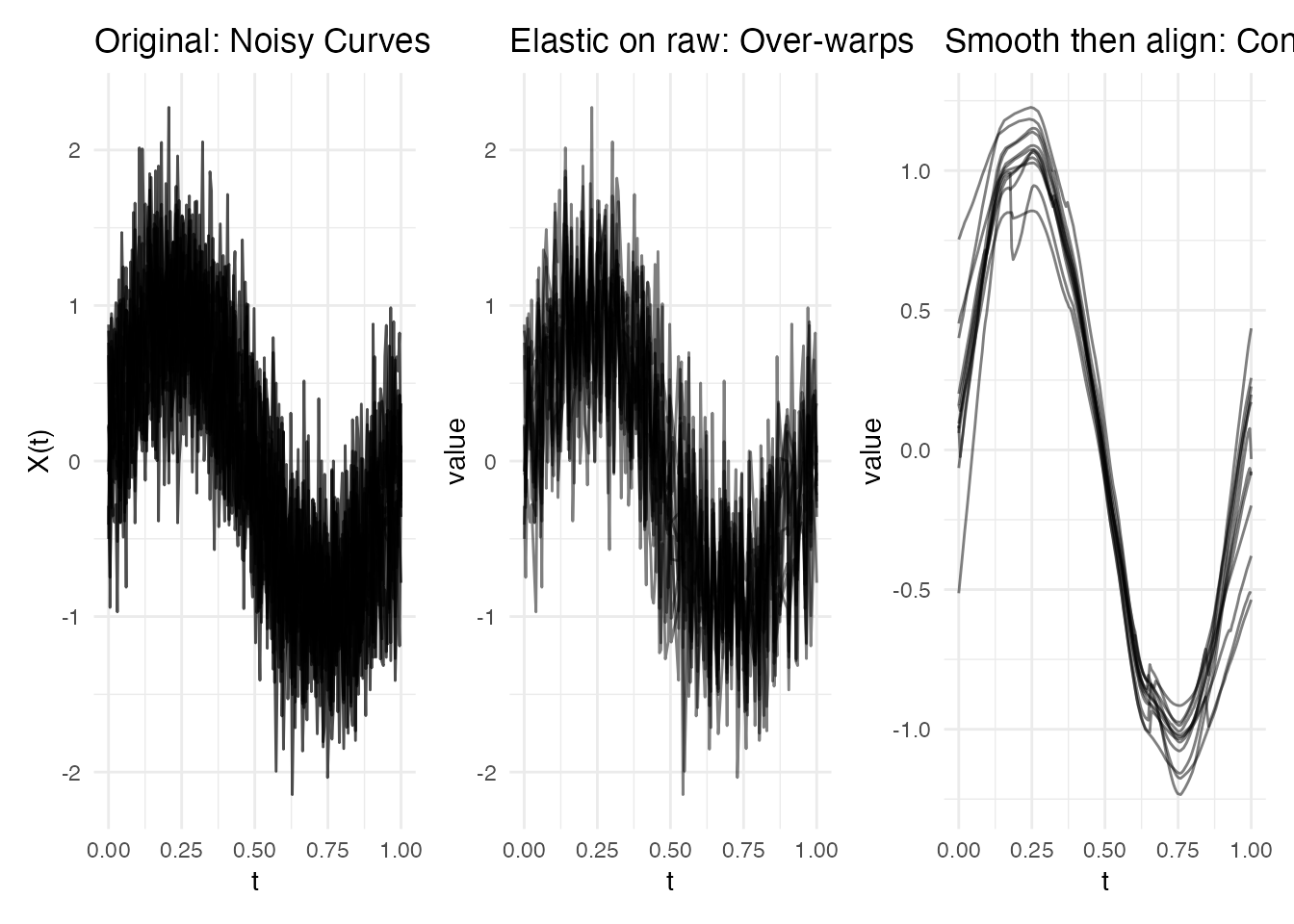
p4 <- plot(ea_noisy_raw, type = "warps") +
labs(title = "Warps (raw): Overly complex")
p5 <- plot(ea_noisy_smooth, type = "warps") +
labs(title = "Warps (smoothed): Reasonable")
p4 + p5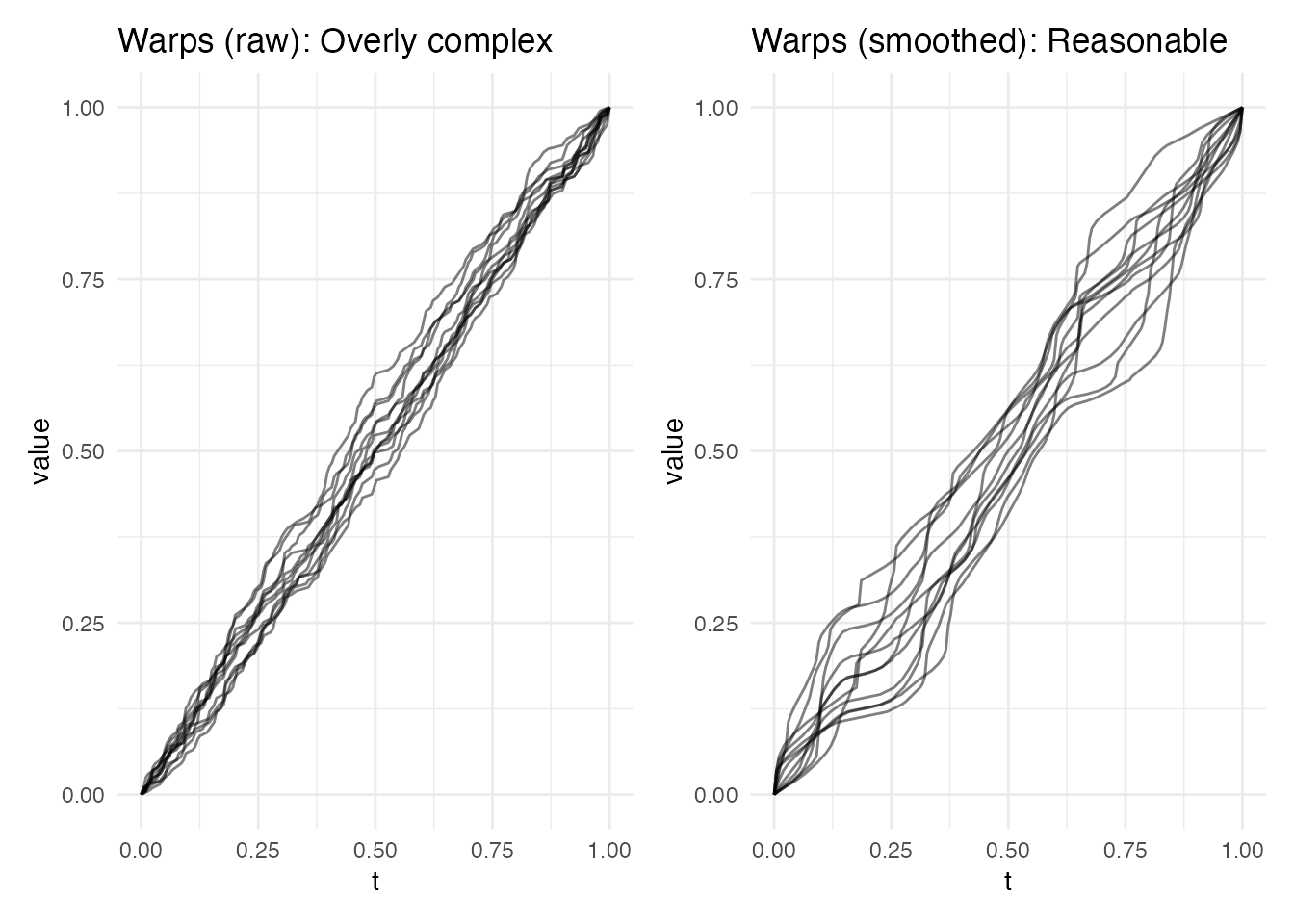
Lesson: Always smooth noisy data before elastic
alignment. The SRSF transform amplifies noise (it involves a
derivative), causing the dynamic programming to match noise spikes.
Pre-smoothing with splines or a kernel smoother prevents over-warping.
Alternatively, use karcher.mean() with its
lambda parameter to penalize warp complexity.
Scenario 4: Landmark Struggles — Variable Number of Features
When curves have different numbers of features, landmark registration
with a fixed expected.count forces all curves into the same
template. This can produce artifacts:
data_var <- matrix(0, n, 200)
for (i in 1:n) {
# Some curves have 1 peak, others have 2
if (i <= n/2) {
pk1 <- runif(1, 0.3, 0.5)
data_var[i, ] <- 1.5 * exp(-300 * (argvals - pk1)^2)
} else {
pk1 <- runif(1, 0.2, 0.35)
pk2 <- runif(1, 0.6, 0.75)
data_var[i, ] <- exp(-300 * (argvals - pk1)^2) + exp(-300 * (argvals - pk2)^2)
}
data_var[i, ] <- data_var[i, ] + 0.02 * rnorm(200)
}
fd_var <- fdata(data_var, argvals = argvals)
# Landmark fails gracefully but results are unreliable
lr_var <- tryCatch(
landmark.register(fd_var, kind = "peak", min.prominence = 0.2, expected.count = 2),
error = function(e) NULL
)
ea_var <- elastic.align(fd_var)
p1 <- plot(fd_var) + labs(title = "Original: Mixed 1-peak & 2-peak")
p2 <- plot(ea_var, type = "aligned") + labs(title = "Elastic: Handles gracefully")
if (!is.null(lr_var)) {
p3 <- plot(lr_var, type = "registered") +
labs(title = "Landmark: Forced 2-peak template")
} else {
p3 <- ggplot() + annotate("text", x = 0.5, y = 0.5,
label = "Landmark failed:\ncannot find 2 peaks in all curves",
size = 4, hjust = 0.5) + theme_void() + labs(title = "Landmark: Failed")
}
p1 + p2 + p3 + plot_layout(ncol = 3)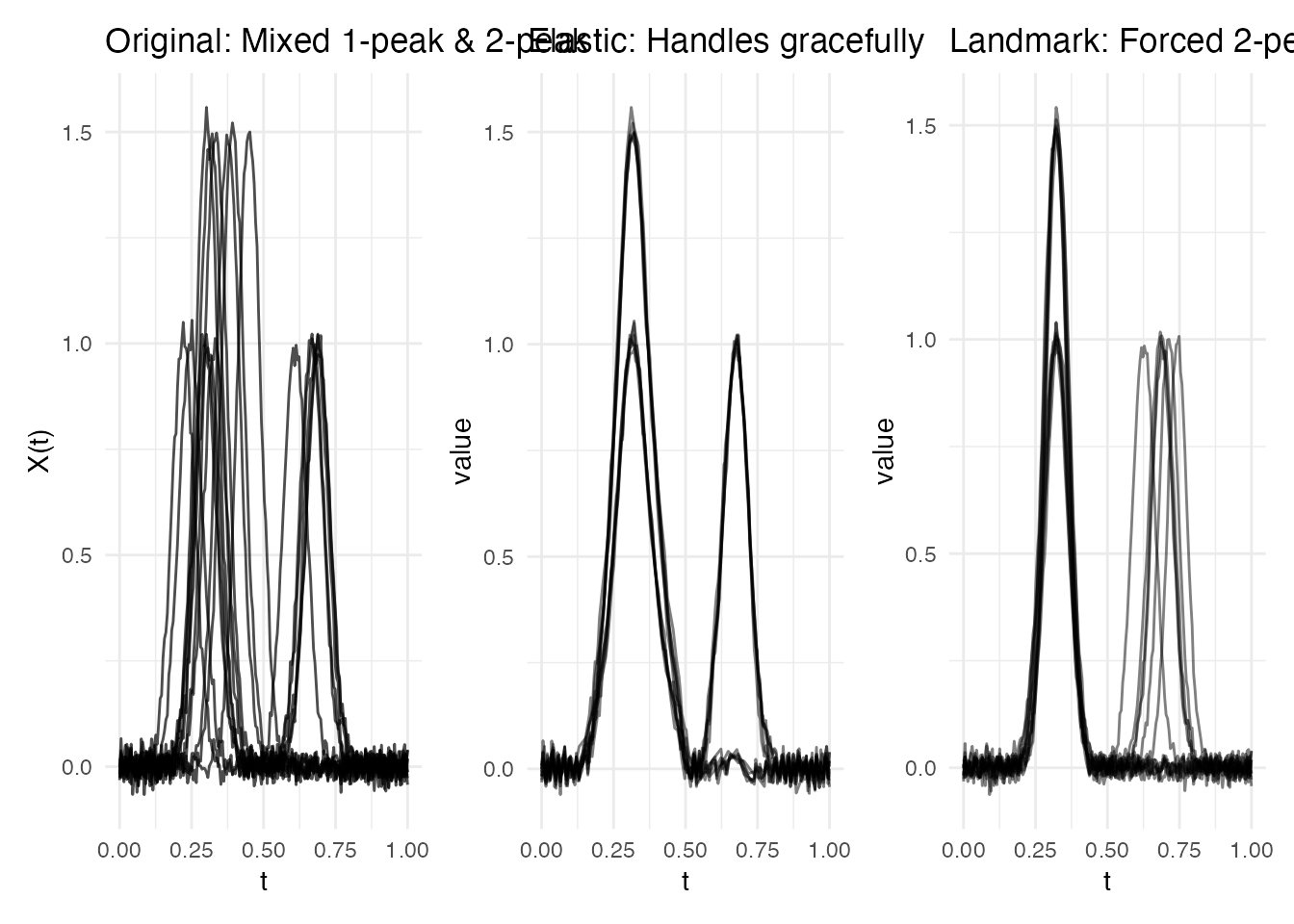
Why elastic wins here: It makes no assumptions about features. When curves have genuinely different shapes, elastic alignment finds the best continuous warping for each pair. Landmark registration requires a consistent feature template across all curves.
Scenario 5: Constrained Excels — Best of Both Worlds
When you have identifiable features AND want smooth warping between them, constrained elastic alignment is the best choice:
# ECG-like data: sharp features + smooth variation between them
data_ecg <- matrix(0, n, 200)
for (i in 1:n) {
r_pos <- runif(1, 0.3, 0.45)
t_pos <- runif(1, 0.55, 0.7)
# R-wave (tall sharp peak)
r_wave <- 2.0 * exp(-800 * (argvals - r_pos)^2)
# T-wave (broader, shorter)
t_wave <- 0.6 * exp(-100 * (argvals - t_pos)^2)
# Baseline wander
baseline <- 0.15 * sin(2 * pi * argvals + runif(1, 0, 2*pi))
data_ecg[i, ] <- r_wave + t_wave + baseline + 0.03 * rnorm(200)
}
fd_ecg <- fdata(data_ecg, argvals = argvals)
ea_ecg <- elastic.align(fd_ecg)
lr_ecg <- landmark.register(fd_ecg, kind = "peak", min.prominence = 0.3,
expected.count = 2)
ec_ecg <- elastic.align.constrained(fd_ecg, kind = "peak",
min.prominence = 0.3,
expected.count = 2)
p1 <- plot(fd_ecg) + labs(title = "Original: ECG-like")
p2 <- plot(ea_ecg, type = "aligned") + labs(title = "Elastic: Good overall")
p3 <- plot(lr_ecg, type = "registered") +
labs(title = "Landmark: Peaks aligned, warps angular")
p4 <- plot(ec_ecg, type = "aligned") +
labs(title = "Constrained: Peaks aligned + smooth warps")
(p1 + p2) / (p3 + p4)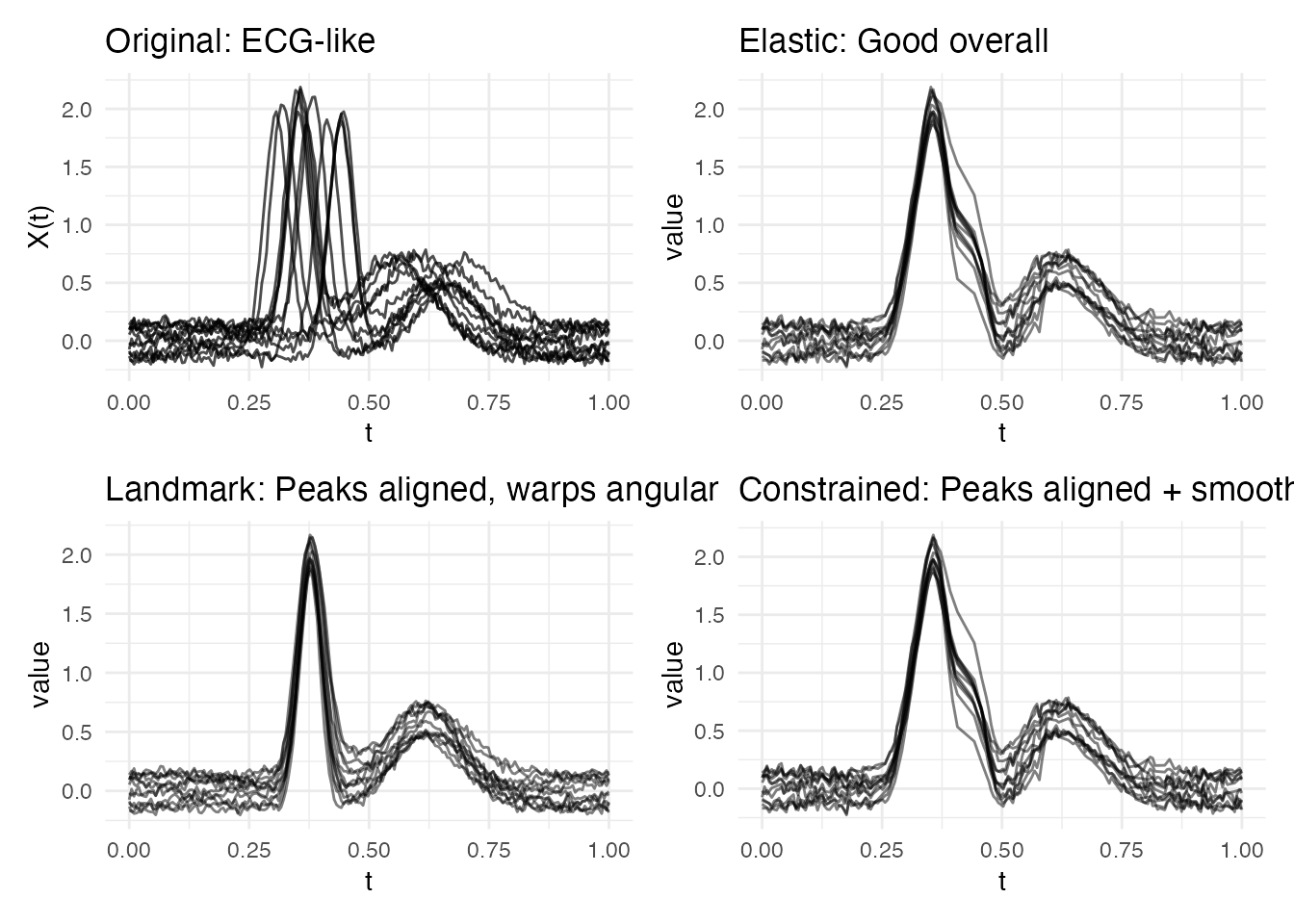
p5 <- plot(ea_ecg, type = "warps") + labs(title = "Elastic warps")
p6 <- plot(lr_ecg, type = "warps") + labs(title = "Landmark warps (piecewise-linear)")
p7 <- plot(ec_ecg, type = "warps") + labs(title = "Constrained warps (smooth + anchored)")
p5 + p6 + p7 + plot_layout(ncol = 3)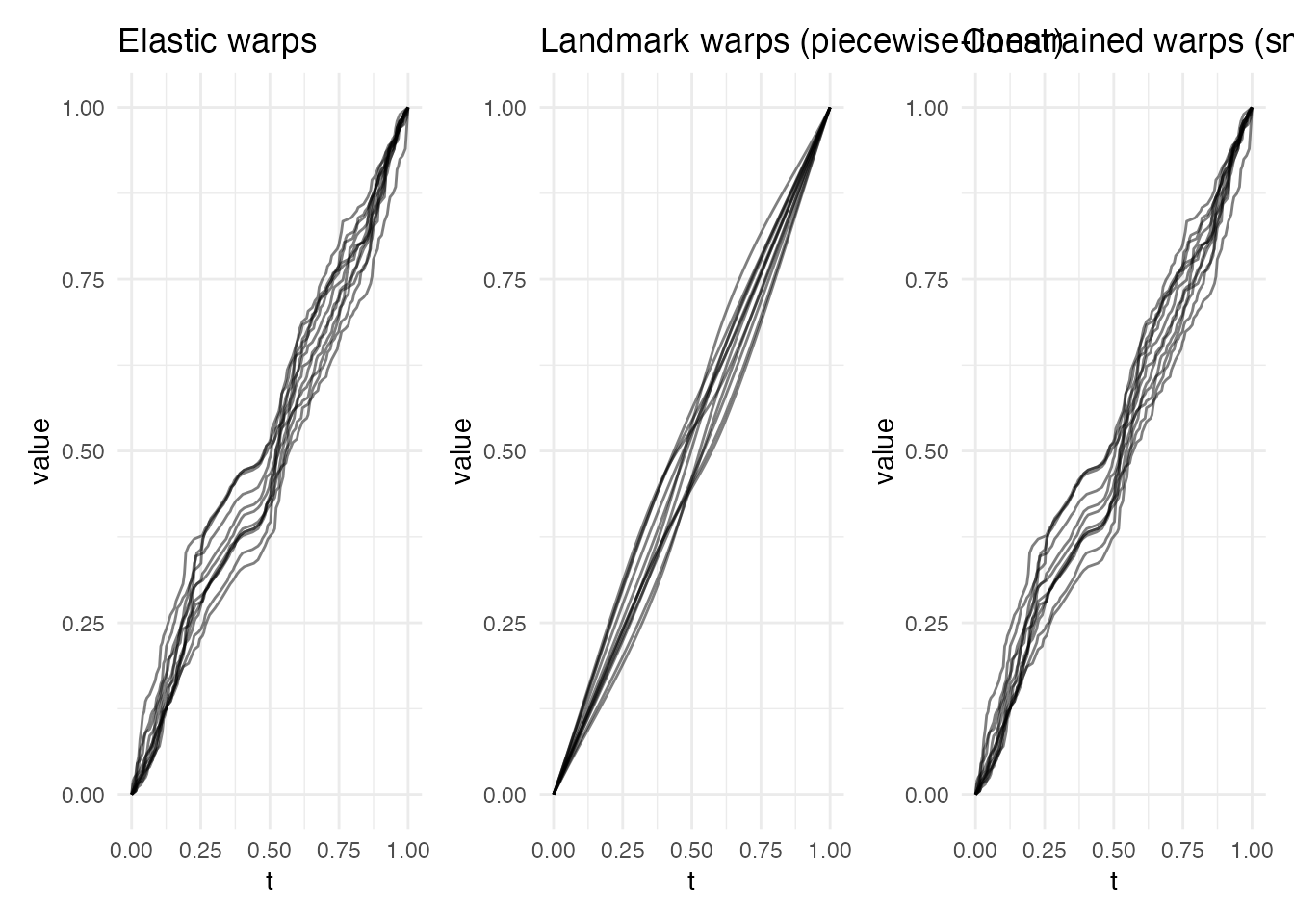
Why constrained wins: The R-wave and T-wave are clinically meaningful features that must correspond correctly (landmark guarantee). But between these features, elastic optimization produces smoother warps than the piecewise-linear interpolation of pure landmark registration.
Scenario Summary
| Scenario | Best method | Why |
|---|---|---|
| Smooth global warping | Elastic | No features to anchor; elastic finds optimal continuous warp |
| Independent feature shifts | Landmark | Guarantees feature correspondence |
| Noisy data | Elastic (with lambda) | Lambda penalty controls over-warping |
| Variable number of features | Elastic | No feature template required |
| Features + smooth variation | Constrained | Combines feature anchoring with smooth optimization |
Decision Guide
Summary Table
| Method | Warping | Automation | Speed | Smoothness | Feature control |
|---|---|---|---|---|---|
| Elastic | Smooth diffeomorphism | Fully automatic | High | None | |
| Landmark | Piecewise-linear | Needs feature type | Low (corners at landmarks) | Full | |
| Constrained | Smooth with anchors | Needs feature type | High (between landmarks) | Partial | |
| TSRVF | (uses elastic) | Fully automatic | + PCA | High | None |
When to Use Each Method
Use elastic alignment when:
- You have no prior knowledge about which features should correspond
- Curves have smooth, continuous timing differences
- You want a principled metric space structure (Fisher-Rao distance)
- You need amplitude-phase decomposition or TSRVF for downstream analysis
Use landmark registration when:
- Features are well-defined and domain-meaningful (ECG waves, spectral peaks)
- You need guaranteed feature correspondence
- Speed is critical (linear in grid size per landmark)
- Curves have complex warping that confuses global DP (e.g., features moving in opposite directions)
Use constrained elastic alignment when:
- You want both smooth warps and feature correspondence
- Some features must align exactly, but the warping between them should be optimized
- You are comparing elastic and landmark results and want a middle ground
Use TSRVF when:
- You need to perform PCA, regression, or classification on aligned data
- You want a Euclidean representation of curve shapes
- You have already computed a Karcher mean and want to extract tangent vectors
Pitfalls
| Method | Common pitfall | Mitigation |
|---|---|---|
| Elastic | Over-warping (pinching) on noisy data | Increase lambda penalty |
| Landmark | Mismatched landmark correspondence | Use expected.count, increase
min.prominence
|
| Constrained | Too few landmarks = barely constrained | Detect multiple landmark types |
| TSRVF | Linearization error for curves far from mean | Check reconstruction quality |
Workflow Example
A typical analysis workflow combining multiple methods:
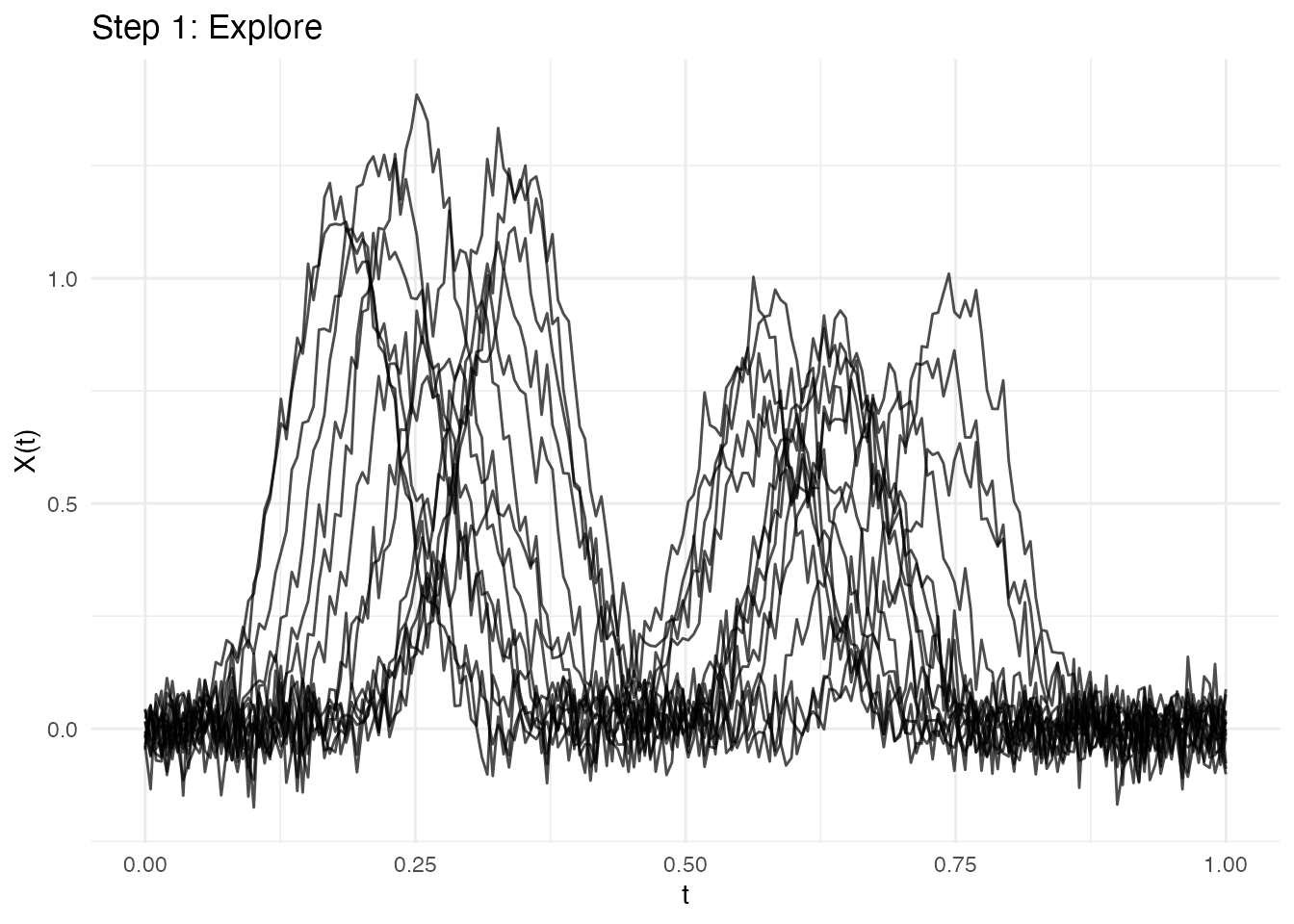
# 2. Detect features to understand structure
lms <- detect.landmarks(fd_feat, kind = "peak", min.prominence = 0.2)
cat("Landmarks per curve:",
paste(sapply(lms, nrow), collapse = ", "), "\n")
#> Landmarks per curve: 3, 2, 3, 5, 2, 2, 3, 2, 4, 2, 4, 2, 2, 3, 3
# 3. Align (choose method based on data)
lr <- landmark.register(fd_feat, kind = "peak", min.prominence = 0.2,
expected.count = 2)
# 4. For downstream analysis, compute Karcher mean + TSRVF
km <- karcher.mean(lr$registered, max.iter = 10)
tv <- tsrvf.from.alignment(km)
# 5. Standard PCA on tangent vectors
pca <- prcomp(tv$tangent_vectors$data)
var_exp <- cumsum(pca$sdev^2 / sum(pca$sdev^2))
cat("Variance explained (first 3 PCs):",
round(var_exp[1:3] * 100, 1), "%\n")
#> Variance explained (first 3 PCs): 26.6 42.6 53.2 %See Also
-
vignette("elastic-alignment", package = "fdars")– elastic alignment framework, quality diagnostics, amplitude-phase decomposition -
vignette("landmark-registration", package = "fdars")– landmark detection, prominence filtering, registration workflow -
vignette("tsrvf", package = "fdars")– TSRVF for linearized elastic analysis, elastic FPCA -
vignette("distance-metrics", package = "fdars")– Soft-DTW, elastic, amplitude, and phase distances