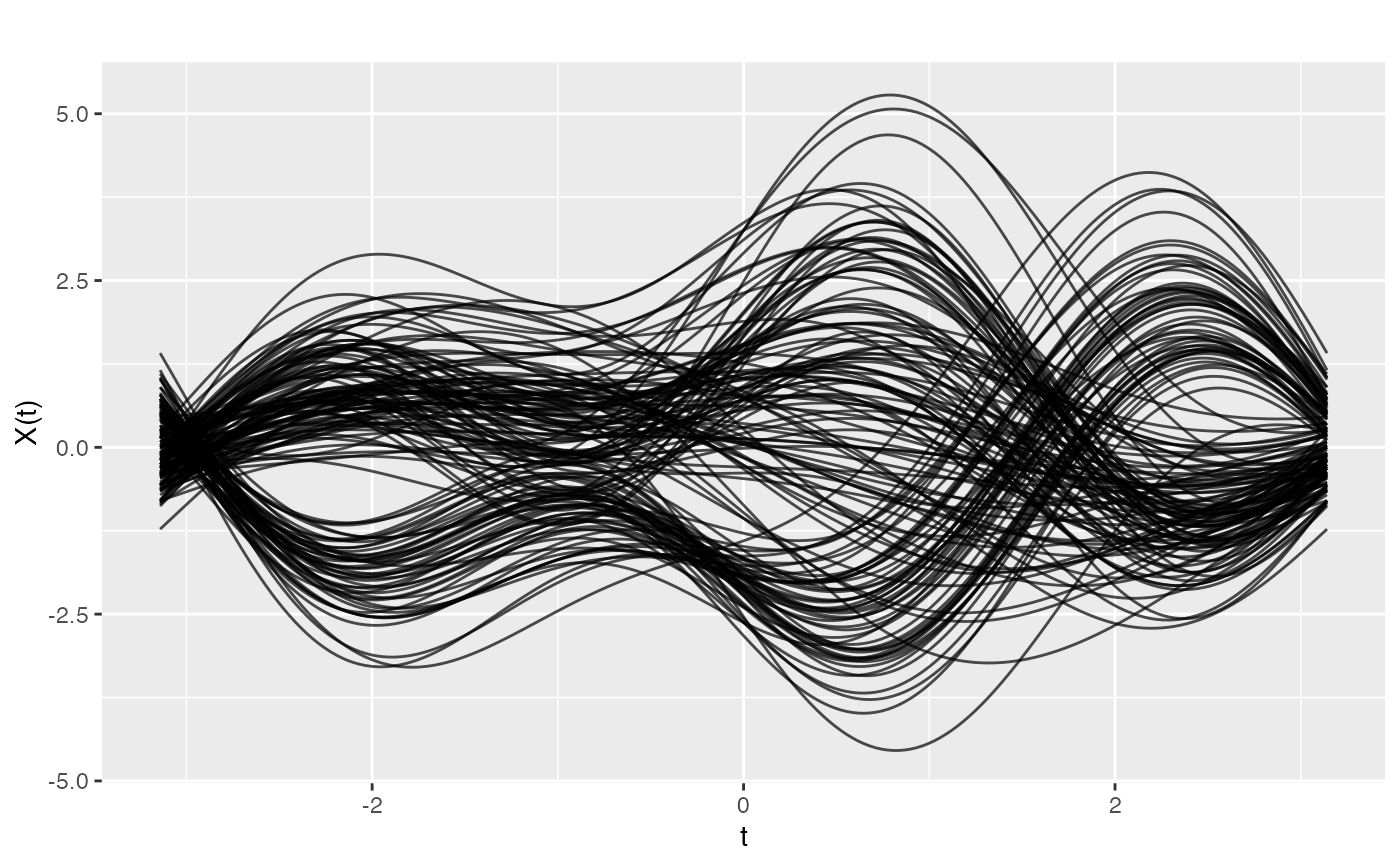
Transforms a multivariate dataset into functional data using the Andrews Fourier expansion. Each row of the input matrix becomes a curve on \([-\pi, \pi]\) defined by $$f_i(t) = x_{i1}/\sqrt{2} + x_{i2}\sin(t) + x_{i3}\cos(t) + x_{i4}\sin(2t) + x_{i5}\cos(2t) + \ldots$$
Arguments
- X
Numeric matrix or data.frame (\(n \times p\)). Rows are observations, columns are variables. Coerced via
as.matrix().- m
Number of grid points on \([-\pi, \pi]\) (default 200).
Value
An fdata object (\(n\) curves, \(m\) grid points) with additional attributes:
andrews_basisThe \(m \times p\) Fourier basis matrix.
andrews_varnamesVariable names (
colnames(X)orpaste0("V", 1:p)ifNULL).
Details
The resulting fdata object carries the Fourier basis matrix and
variable names as attributes, enabling andrews_loadings() to project
FPCA eigenfunctions back to the original variable space.