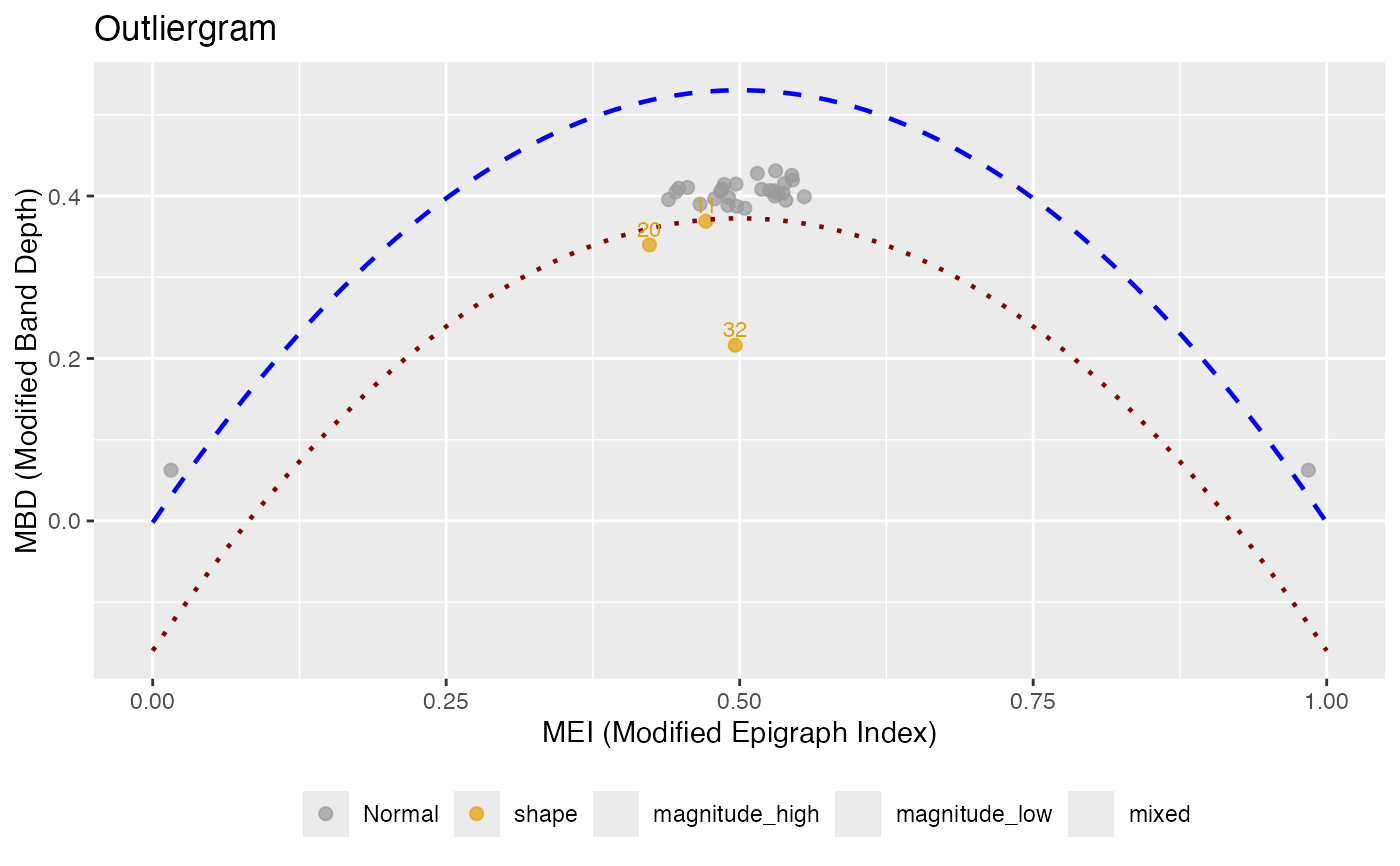
Creates an outliergram plot that displays MEI (Modified Epigraph Index) versus MBD (Modified Band Depth) for outlier detection. Points below the parabolic boundary are identified as outliers, and each outlier is classified by type.
Value
An object of class 'outliergram' with components:
- fdataobj
The input functional data
- mei
MEI values for each curve
- mbd
MBD values for each curve
- outliers
Indices of detected outliers
- outlier_type
Character vector of outlier types ("shape") for each detected outlier
- n_outliers
Number of outliers detected
- factor
The factor used for threshold adjustment
- parabola
Coefficients of the parabolic boundary (a0, a1, a2)
- threshold
The boxplot-fence threshold for distance below the parabola
- dist_to_parabola
Vertical distance below the parabola for each curve (positive values indicate the point is below the parabola)
Details
The outliergram plots MEI on the x-axis versus MBD on the y-axis. For a sample of size \(n\), the theoretical relationship is bounded by the finite-sample parabola (Arribas-Gil & Romo, 2014, Proposition 1): $$MBD \le a_0 + a_1 \cdot MEI + a_2 \cdot MEI^2$$ where \(a_0 = -2/(n(n-1))\), \(a_1 = 2(n+1)/(n-1)\), \(a_2 = -2(n+1)/(n-1)\).
Shape outliers are detected using a boxplot fence on the vertical distances below the parabola: a curve is flagged when its distance exceeds \(Q_3 + \mathrm{factor} \times IQR\).
References
Arribas-Gil, A. and Romo, J. (2014). Shape outlier detection and visualization for functional data: the outliergram. Biostatistics, 15(4), 603-619.
See also
depth for depth computation, magnitudeshape for
an alternative outlier visualization.
Examples
# Create functional data with different outlier types
set.seed(42)
t <- seq(0, 1, length.out = 50)
X <- matrix(0, 32, 50)
for (i in 1:29) X[i, ] <- sin(2 * pi * t) + rnorm(50, sd = 0.2)
X[30, ] <- sin(2 * pi * t) + 2 # magnitude outlier (high)
X[31, ] <- sin(2 * pi * t) - 2 # magnitude outlier (low)
X[32, ] <- sin(4 * pi * t) # shape outlier
fd <- fdata(X, argvals = t)
# Create outliergram
og <- outliergram(fd)
print(og)
#> Outliergram
#> ===========
#> Number of curves: 32
#> Outliers detected: 3
#>
#> Outlier types:
#> Shape: 3
#>
#> Outlier details:
#> Index 11 : shape
#> Index 20 : shape
#> Index 32 : shape
#>
#> Outlier p-values: 0.1212, 0.0909, 0.0606
#>
#> Parameters:
#> Factor: 1.5
plot(og, color_by_type = TRUE)