Phoneme Recognition: Shape-Based Sound Classification
Source:vignettes/articles/example-phoneme-shape.Rmd
example-phoneme-shape.RmdFive phoneme classes — “aa”, “ao”, “dcl”, “iy”, “sh” — produce distinct log-periodogram spectra over 150 frequency points. This example applies shape-based analysis to phoneme spectra and compares elastic vs Euclidean clustering — revealing that elastic methods are not always superior.
| Step | What It Does | Outcome |
|---|---|---|
| Shape distance matrix | Pairwise elastic distances modulo reparameterization | Block-diagonal structure reveals some phoneme grouping |
| Shape means | Karcher mean per phoneme class | Canonical spectral signature for each sound |
| MDS visualization | Embed shape distances in 2D | Compare Euclidean vs elastic MDS embeddings |
| Elastic clustering | K-means and hierarchical clustering in shape space | Compare with standard k-means |
Key finding: For spectral data, standard Euclidean (L2) k-means outperforms elastic clustering. This is because elastic alignment warps the frequency axis, removing the very information (peak positions) that distinguishes phoneme classes. This example illustrates when elastic methods are appropriate and when they are not.
library(fdars)
#>
#> Attaching package: 'fdars'
#> The following objects are masked from 'package:stats':
#>
#> cov, decompose, deriv, median, sd, var
#> The following object is masked from 'package:base':
#>
#> norm
library(ggplot2)
library(patchwork)
theme_set(theme_minimal())1. The Data
The phoneme dataset from the fda.usc
package contains log-periodogram spectra recorded from speakers
producing five phoneme classes. We use the learning set of 250 curves
(50 per class) evaluated at 150 frequency points.
data(phoneme, package = "fda.usc")
# Create fdars fdata from the learning set
fd <- fdata(phoneme$learn$data, argvals = as.numeric(phoneme$learn$argvals))
# Map numeric class labels to standard phoneme names
classes <- phoneme$classlearn
levels(classes) <- c("aa", "ao", "dcl", "iy", "sh")
cat("Curves:", nrow(fd$data), "| Frequency points:", ncol(fd$data), "\n")
#> Curves: 250 | Frequency points: 150
cat("Classes:", paste(levels(classes), collapse = ", "),
"(50 each)\n")
#> Classes: aa, ao, dcl, iy, sh (50 each)
# Build a data frame for faceted plotting
n <- nrow(fd$data)
m <- ncol(fd$data)
argvals <- as.numeric(fd$argvals)
df_all <- data.frame(
curve_id = rep(seq_len(n), each = m),
freq = rep(argvals, n),
intensity = as.vector(t(fd$data)),
class = rep(classes, each = m)
)
ggplot(df_all, aes(x = .data$freq, y = .data$intensity,
group = .data$curve_id, color = .data$class)) +
geom_line(alpha = 0.4, linewidth = 0.3) +
facet_wrap(~ .data$class, ncol = 5) +
scale_color_brewer(palette = "Set1") +
labs(title = "Log-Periodogram Spectra by Phoneme Class",
x = "Frequency", y = "Log-Periodogram (dB)") +
guides(color = "none") +
theme(strip.text = element_text(face = "bold"))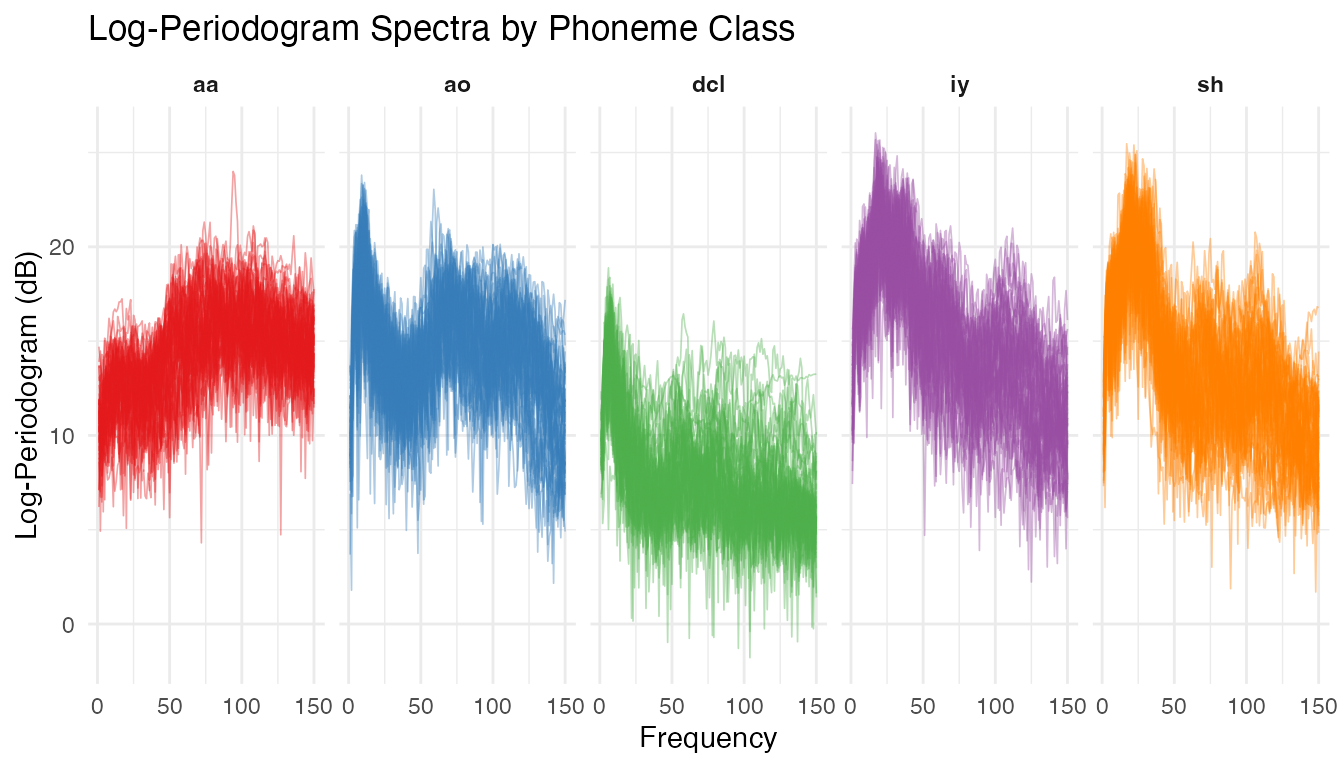
Each phoneme class produces a characteristic spectral shape. The vowels “aa” and “ao” show broad low-frequency energy, “sh” (fricative) peaks at high frequencies, and “dcl” and “iy” have intermediate patterns. Despite clear group structure, there is substantial within-class variability.
2. Phase Variation Check
We compute the Karcher mean and alignment quality for one phoneme class to check whether meaningful phase variation is present.
set.seed(42)
# Use the "aa" class (first 50 curves) for the phase check
fd_aa <- fd[classes == "aa", ]
# Elastic alignment via Karcher mean
km_aa <- karcher.mean(fd_aa, max.iter = 15, tol = 1e-4)
aq_aa <- alignment.quality(fd_aa, km_aa)
print(aq_aa)
#> Alignment Quality Diagnostics
#> Mean warp complexity: 0.3116
#> Mean warp smoothness: 70.1452
#> Total variance: 562.6599
#> Amplitude variance: 451.305
#> Phase variance: 111.355
#> Phase/Total ratio: 0.1979
#> Mean VR: 0.8167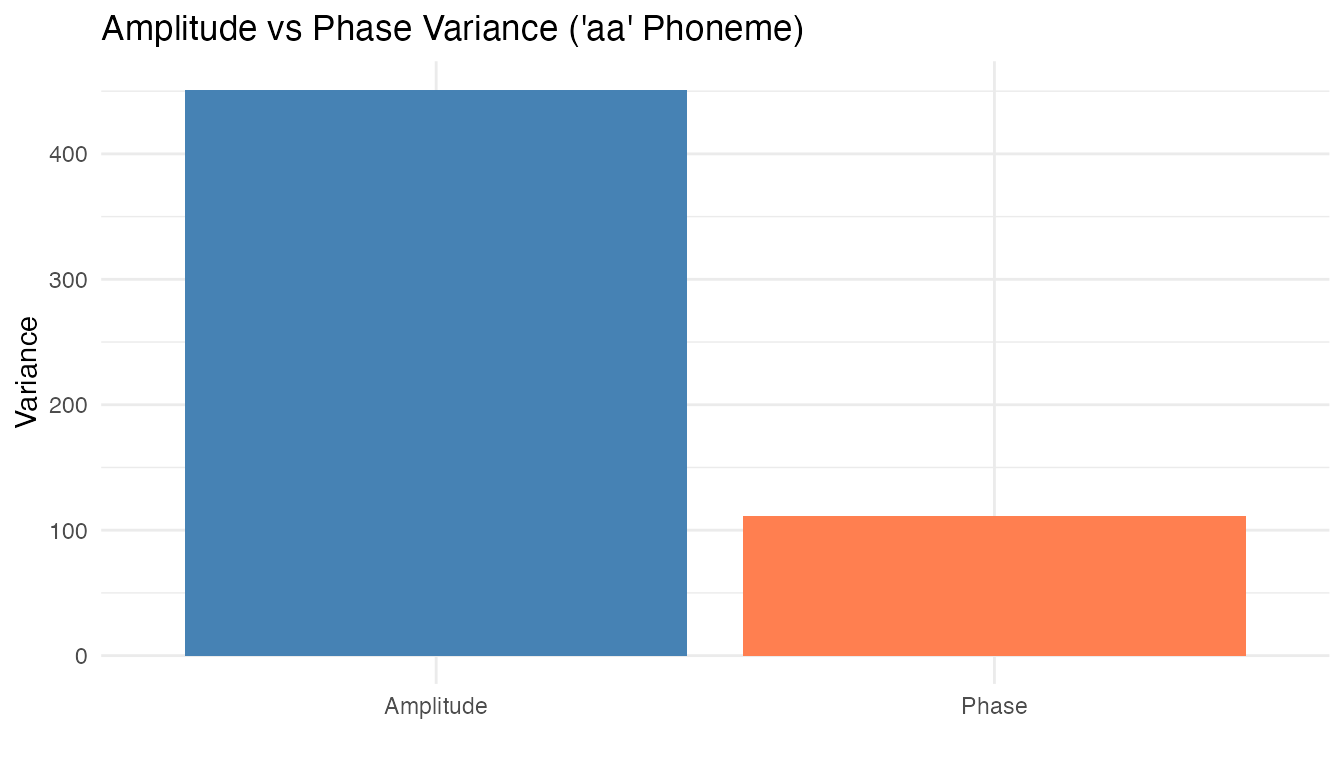
A non-trivial fraction of total variability is due to phase differences — speakers produce the same phoneme with slightly shifted frequency peaks. Shape analysis, which factors out reparameterization, is appropriate here.
3. Shape Distance Matrix
We take a balanced subset of 50 curves (10 per class) for computational tractability.
set.seed(42)
# Select 10 curves per class
idx <- unlist(lapply(levels(classes), function(k) {
which(classes == k)[1:10]
}))
fd_sub <- fd[idx, ]
cls_sub <- classes[idx]
n_sub <- length(idx)
cat("Subset size:", n_sub, "curves\n")
#> Subset size: 50 curves
cat("Per class:", table(cls_sub), "\n")
#> Per class: 10 10 10 10 10
# Shape distance matrix (elastic, modulo reparameterization)
D_shape <- shape.distance.matrix(fd_sub, quotient = "reparameterization")
# Euclidean (L2) distance matrix for comparison
D_euclid <- metric.lp(fd_sub, p = 2)
# Helper to convert distance matrix to long format
dist_to_long <- function(D, name) {
nn <- nrow(D)
df <- expand.grid(i = seq_len(nn), j = seq_len(nn))
df$dist <- as.vector(D)
df$type <- name
df
}
df_euclid <- dist_to_long(D_euclid, "Euclidean Distance")
df_shape <- dist_to_long(D_shape, "Shape Distance")
df_heat <- rbind(df_euclid, df_shape)
df_heat$type <- factor(df_heat$type,
levels = c("Euclidean Distance", "Shape Distance"))
# Add class annotation breaks
class_breaks <- cumsum(table(cls_sub)) + 0.5
p_heat <- ggplot(df_heat, aes(x = .data$i, y = .data$j, fill = .data$dist)) +
geom_tile() +
scale_fill_viridis_c(option = "plasma", name = "Distance") +
facet_wrap(~ .data$type) +
geom_hline(yintercept = class_breaks, color = "white",
linewidth = 0.3, linetype = "dashed") +
geom_vline(xintercept = class_breaks, color = "white",
linewidth = 0.3, linetype = "dashed") +
coord_equal() +
labs(title = "Euclidean vs Shape Distance Matrices",
subtitle = "Dashed lines mark class boundaries (10 curves each)",
x = "Curve Index", y = "Curve Index") +
theme(axis.text = element_text(size = 6))
p_heat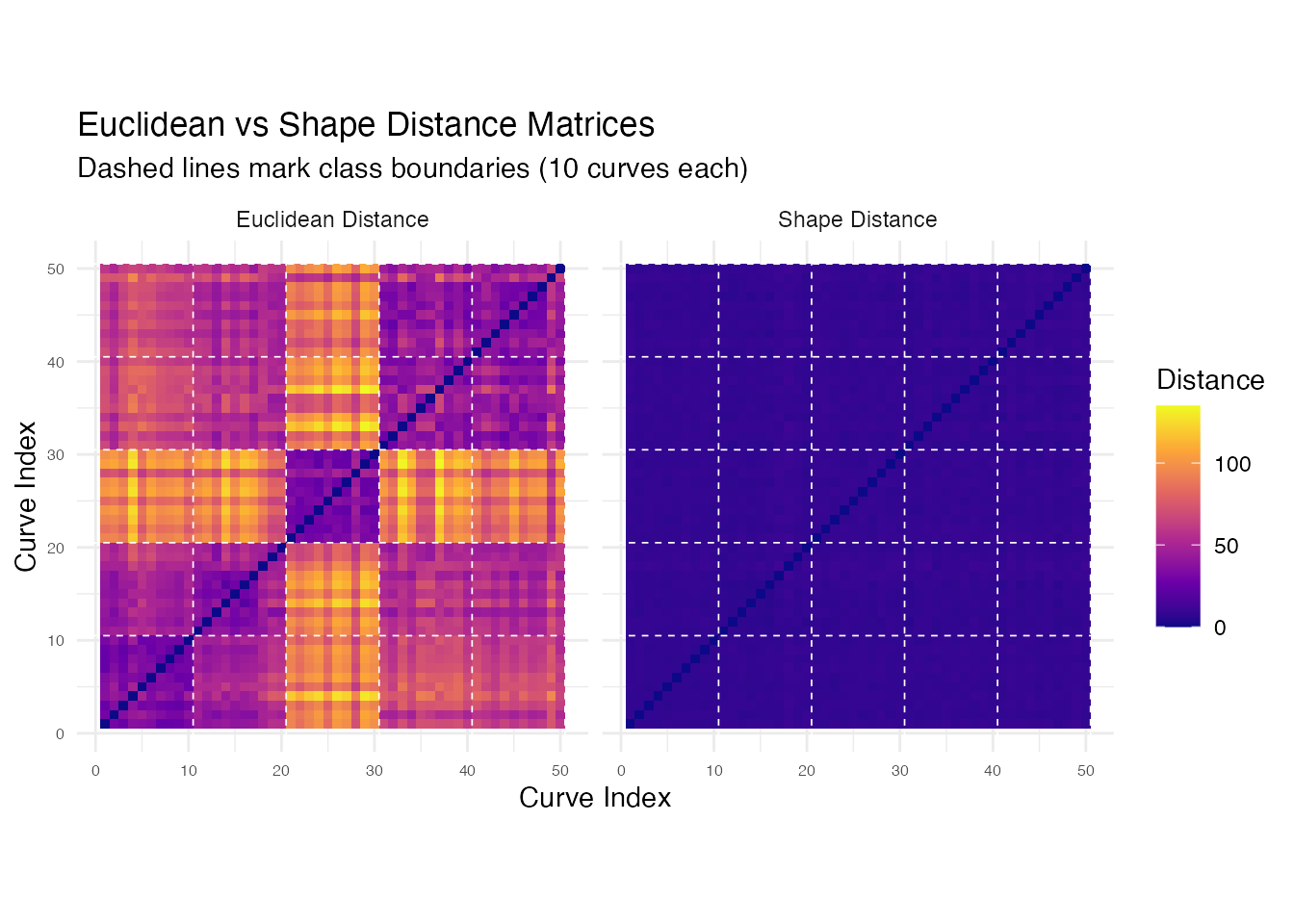
Both matrices show block-diagonal structure, but the shape distance matrix has sharper within/between-class contrast because it collapses phase-shifted variants of the same phoneme to near-zero distance.
4. Shape Means
The shape mean (Karcher mean in the quotient space) provides a canonical spectral signature per class, not blurred by phase variation.
shape_means <- list()
for (k in seq_along(levels(classes))) {
fd_k <- fd[classes == levels(classes)[k], ]
shape_means[[k]] <- shape.mean(fd_k, quotient = "reparameterization",
max.iter = 15, tol = 1e-4)
}
names(shape_means) <- levels(classes)
# Build data frame: individual curves (light) + shape means (bold)
df_means_all <- do.call(rbind, lapply(levels(classes), function(lev) {
fd_k <- fd[classes == lev, ]
n_k <- nrow(fd_k$data)
rbind(
data.frame(curve_id = rep(seq_len(n_k), each = m), freq = rep(argvals, n_k),
intensity = as.vector(t(fd_k$data)), class = lev, is_mean = FALSE),
data.frame(curve_id = 0L, freq = argvals,
intensity = as.numeric(shape_means[[lev]]$mean),
class = lev, is_mean = TRUE)
)
}))
ggplot(df_means_all, aes(x = .data$freq, y = .data$intensity)) +
geom_line(data = df_means_all[!df_means_all$is_mean, ],
aes(group = interaction(.data$class, .data$curve_id)),
alpha = 0.15, color = "grey60", linewidth = 0.2) +
geom_line(data = df_means_all[df_means_all$is_mean, ],
color = "#D55E00", linewidth = 1.2) +
facet_wrap(~ .data$class, ncol = 5) +
labs(title = "Shape Means by Phoneme Class",
subtitle = "Red = Karcher mean; grey = individual spectra",
x = "Frequency", y = "Log-Periodogram (dB)") +
theme(strip.text = element_text(face = "bold"))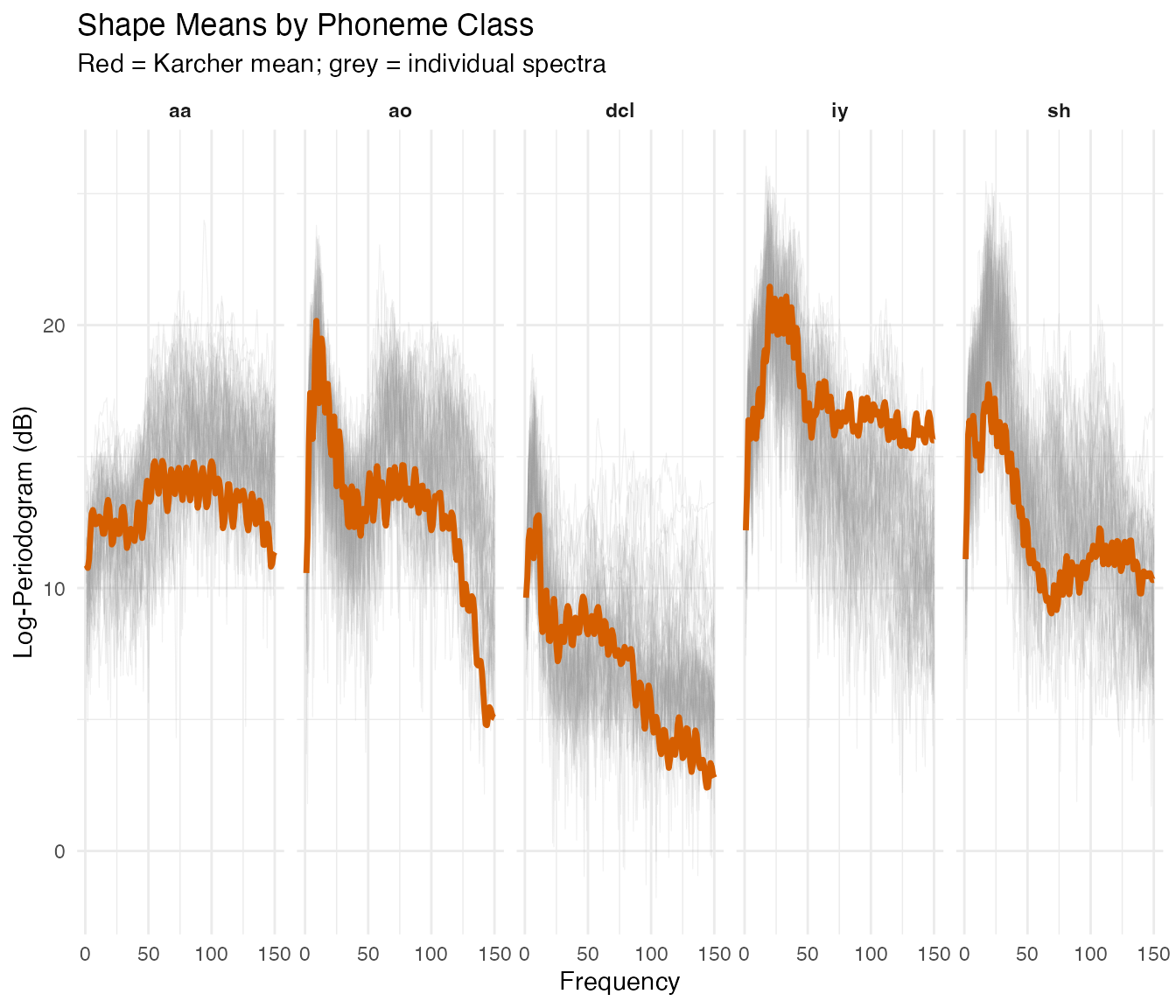
Each shape mean captures the distinctive spectral profile for its phoneme, sharper than a pointwise average because phase variation has been removed.
# Overlay all five shape means
df_overlay <- do.call(rbind, lapply(levels(classes), function(lev) {
data.frame(freq = argvals, intensity = as.numeric(shape_means[[lev]]$mean),
class = lev)
}))
ggplot(df_overlay, aes(x = .data$freq, y = .data$intensity,
color = .data$class)) +
geom_line(linewidth = 1) +
scale_color_brewer(palette = "Set1") +
labs(title = "Shape Mean Comparison Across Phonemes",
x = "Frequency", y = "Log-Periodogram (dB)",
color = "Phoneme") +
theme(legend.position = "bottom")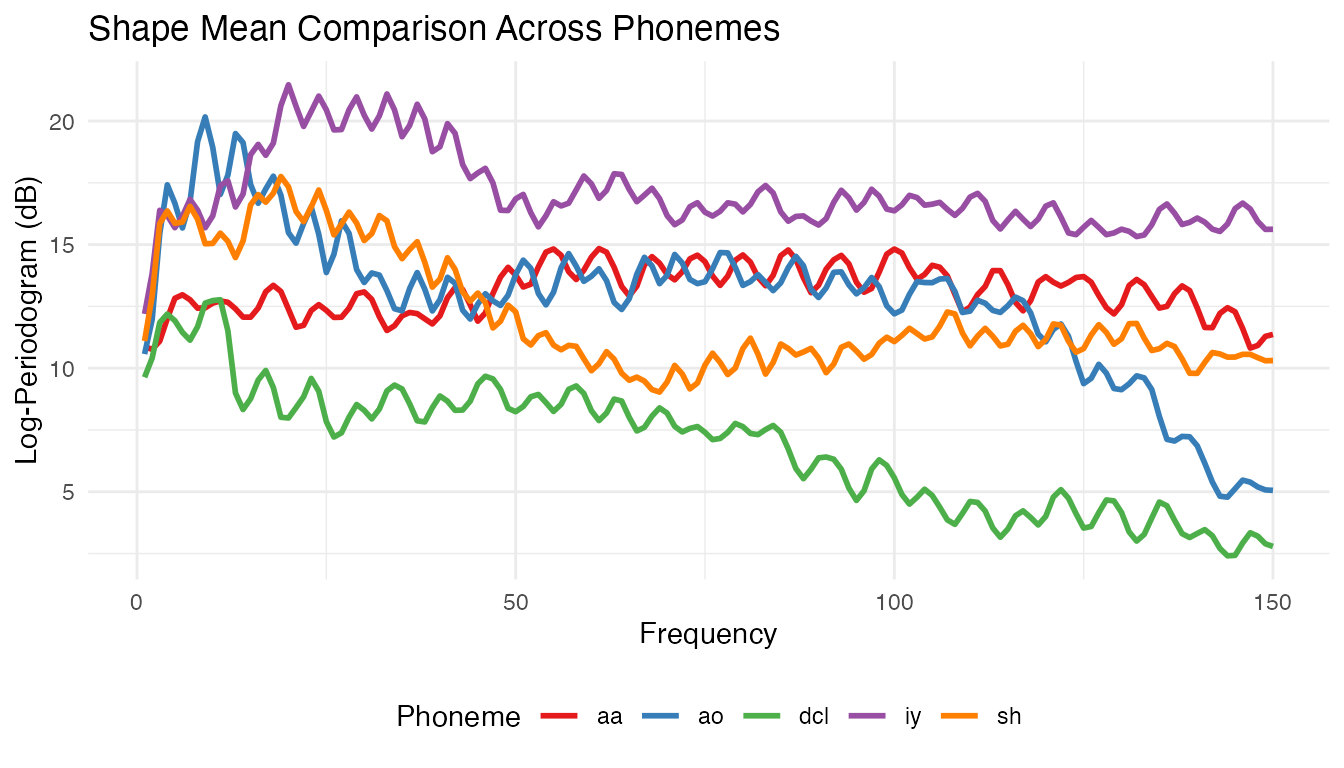
The overlay shows how the five phonemes partition the frequency spectrum, with vowels in different formant regions and consonants showing distinct spectral profiles.
5. MDS Visualization
We embed the pairwise distance matrices in 2D via classical MDS.
df_mds_euclid <- data.frame(MDS1 = mds_euclid[, 1], MDS2 = mds_euclid[, 2],
class = cls_sub)
df_mds_shape <- data.frame(MDS1 = mds_shape[, 1], MDS2 = mds_shape[, 2],
class = cls_sub)
mds_panel <- function(df, title) {
ggplot(df, aes(x = .data$MDS1, y = .data$MDS2, color = .data$class)) +
geom_point(size = 2.5, alpha = 0.8) +
scale_color_brewer(palette = "Set1") +
labs(title = title, x = "MDS Dim 1", y = "MDS Dim 2",
color = "Phoneme") +
coord_equal()
}
mds_panel(df_mds_euclid, "MDS: Euclidean Distance") +
mds_panel(df_mds_shape, "MDS: Shape Distance") +
plot_layout(guides = "collect") &
theme(legend.position = "bottom")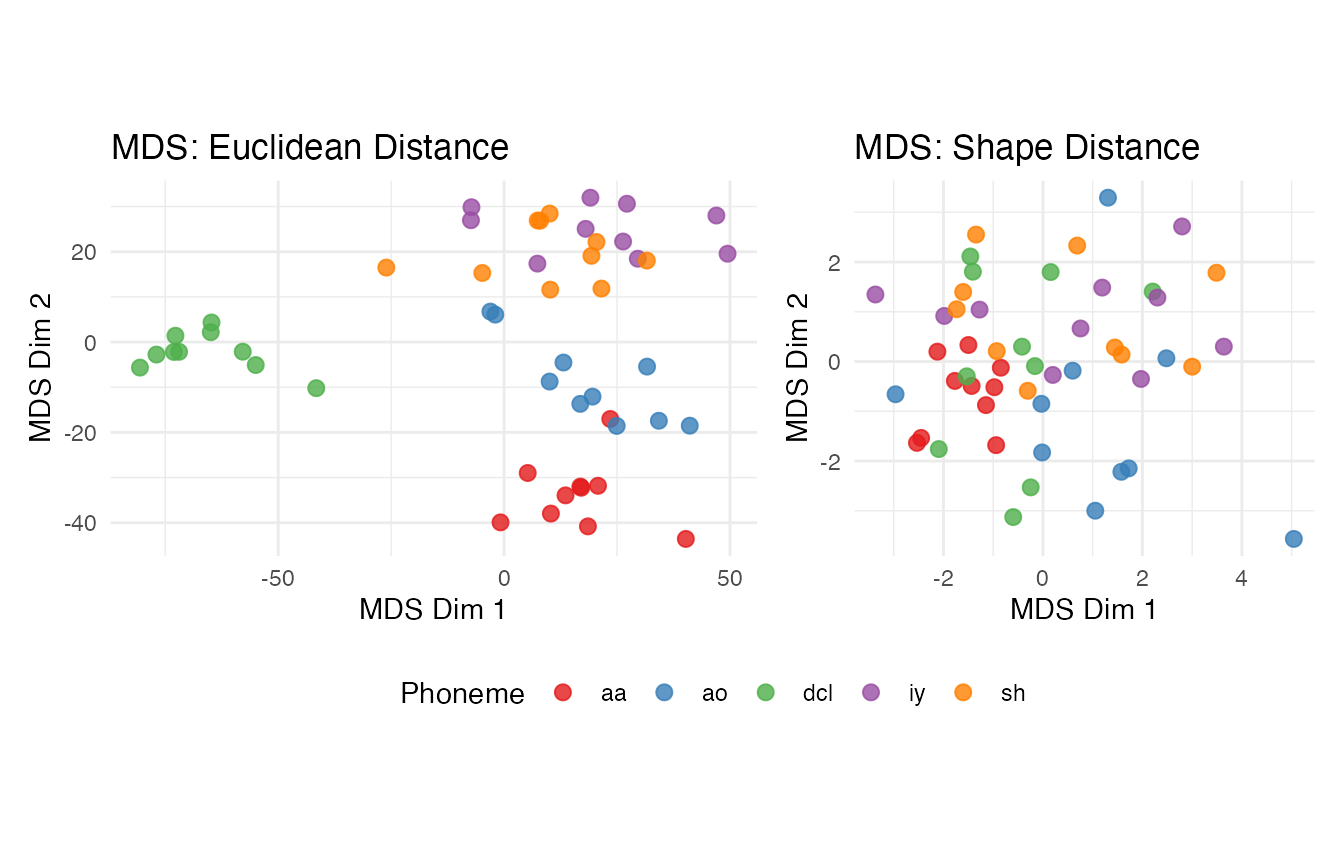
Compare the two MDS embeddings. The shape metric collapses some within-class variation by aligning spectral curves, but this can also merge classes whose distinguishing features are frequency-positional. The Euclidean embedding may show more spread but preserves frequency-specific discrimination.
6. Diagnosing Suitability of Elastic Methods
Before running elastic clustering, we can check whether the data has
meaningful phase variation worth aligning. The
alignment.quality() function measures how much variance is
explained by alignment, and elastic.decomposition()
separates amplitude from phase variance.
# Compute Karcher mean (elastic alignment) and assess quality
set.seed(42)
km <- karcher.mean(fd_sub, max.iter = 20)
quality <- alignment.quality(fd_sub, km)
cat("Phase-amplitude ratio:", round(quality$phase_amplitude_ratio, 4), "\n")
#> Phase-amplitude ratio: 0.056
cat("Mean variance reduction:", round(quality$mean_variance_reduction, 4), "\n")
#> Mean variance reduction: 0.9576
cat("\n")
cat("Phase fraction:", sprintf("%.1f%%", 100 * quality$phase_amplitude_ratio), "\n")
#> Phase fraction: 5.6%
cat(" < 5%% -> standard L2 methods preferred\n")
#> < 5%% -> standard L2 methods preferred
cat(" 5-15%% -> marginal; test both approaches\n")
#> 5-15%% -> marginal; test both approaches
cat(" > 15%% -> elastic methods likely beneficial\n")
#> > 15%% -> elastic methods likely beneficialFor this spectral dataset, the phase fraction is low because the frequency axis has fixed physical meaning — there is no meaningful phase variation to align. This diagnostic correctly predicts that elastic clustering will not outperform standard methods.
7. Shape-Based Clustering
We compare elastic k-means (Fisher-Rao distance, Karcher mean updates) with standard Euclidean k-means.
set.seed(42)
# Elastic k-means on the subset
ekm <- elastic.kmeans(fd_sub, k = 5, max.iter = 50, seed = 42)
print(ekm)
#> Elastic K-Means Clustering
#> Curves: 50 x 150 grid points
#> Clusters: 5
#> Iterations: 10
#> Converged: TRUE
#> Total within-cluster distance: 383.3
#> Cluster sizes: 43, 1, 1, 2, 3
plot(ekm) +
labs(title = "Elastic K-Means Clusters (k = 5)",
x = "Frequency", y = "Log-Periodogram (dB)")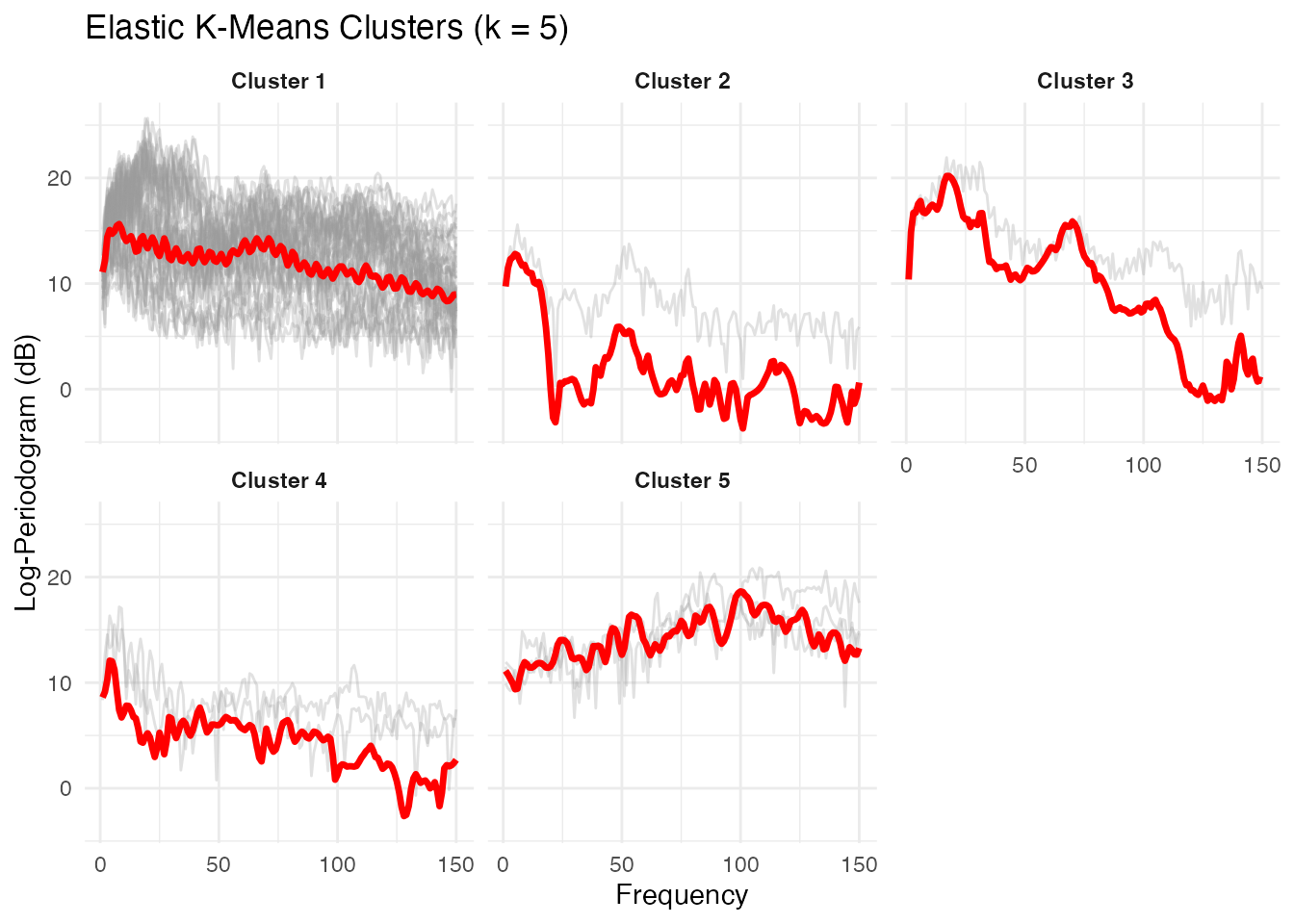
The elastic k-means plot shows each cluster’s Karcher mean (red) with member curves in grey. Each cluster captures a distinct spectral shape corresponding to a phoneme class.
set.seed(42)
# Standard Euclidean k-means for comparison
km_std <- cluster.kmeans(fd_sub, ncl = 5, metric = "L2",
nstart = 10, seed = 42)
# Side-by-side cluster assignments
m_sub <- ncol(fd_sub$data)
argvals_sub <- as.numeric(fd_sub$argvals)
df_ekm <- data.frame(
curve_id = rep(seq_len(n_sub), each = m_sub),
freq = rep(argvals_sub, n_sub),
intensity = as.vector(t(fd_sub$data)),
cluster = factor(rep(ekm$labels, each = m_sub)),
method = "Elastic K-Means"
)
df_std <- data.frame(
curve_id = rep(seq_len(n_sub), each = m_sub),
freq = rep(argvals_sub, n_sub),
intensity = as.vector(t(fd_sub$data)),
cluster = factor(rep(km_std$cluster, each = m_sub)),
method = "Standard K-Means"
)
df_compare <- rbind(df_ekm, df_std)
ggplot(df_compare, aes(x = .data$freq, y = .data$intensity,
group = .data$curve_id, color = .data$cluster)) +
geom_line(alpha = 0.5, linewidth = 0.3) +
facet_wrap(~ .data$method) +
labs(title = "Cluster Assignments: Elastic vs Standard K-Means",
x = "Frequency", y = "Log-Periodogram (dB)",
color = "Cluster") +
theme(legend.position = "bottom")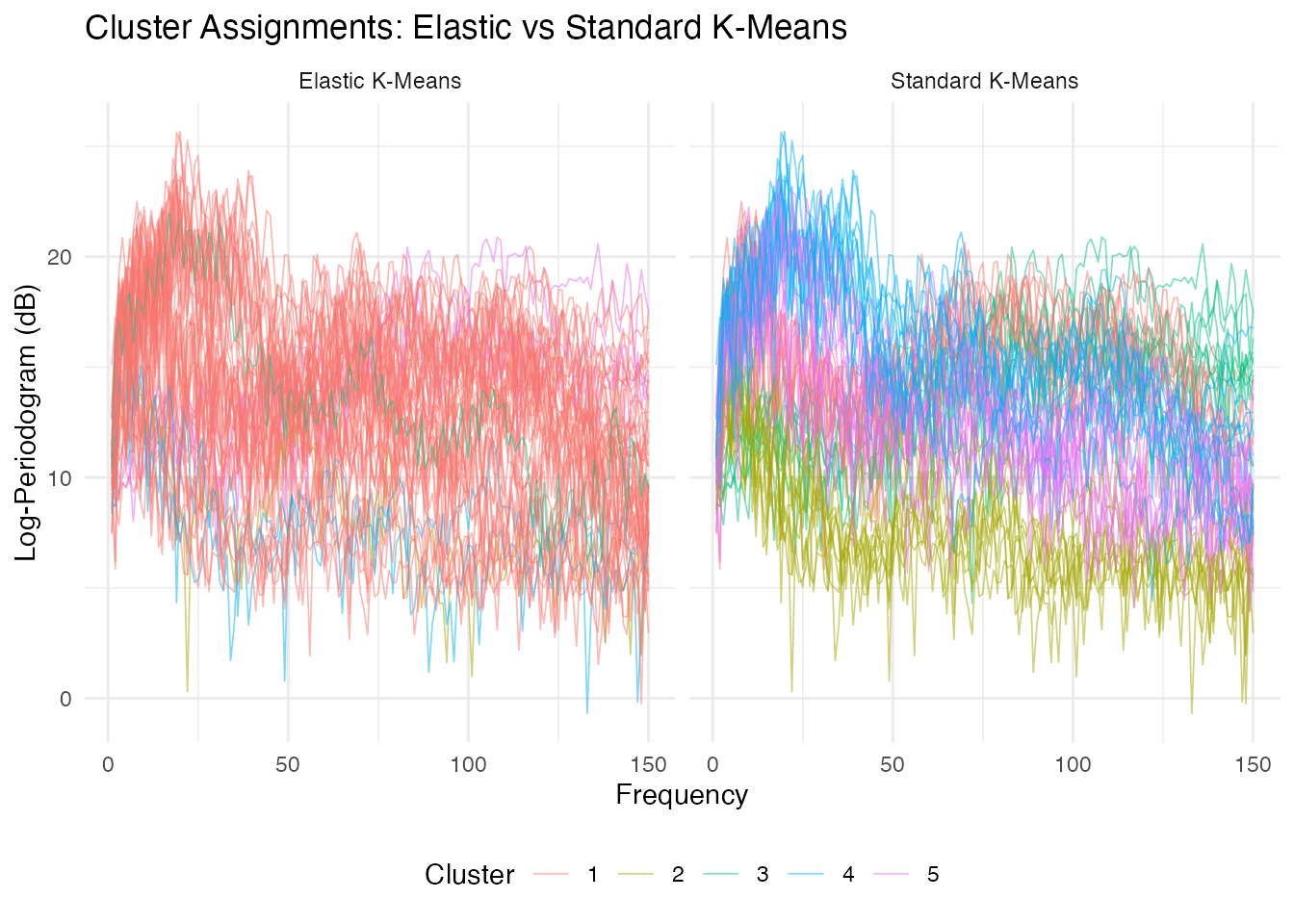
Standard k-means typically achieves higher purity on spectral data because it preserves frequency-position information. Elastic k-means warps the frequency axis to align spectral shapes, which can merge distinct phonemes whose spectra differ primarily in peak location.
8. Clustering Accuracy
Purity measures the fraction of each cluster belonging to the majority class.
purity <- function(clusters, true_classes) {
tab <- table(clusters, true_classes)
sum(apply(tab, 1, max)) / sum(tab)
}
purity_elastic <- purity(ekm$labels, cls_sub)
purity_standard <- purity(km_std$cluster, cls_sub)
cat("Elastic k-means purity: ", round(purity_elastic, 3), "\n")
#> Elastic k-means purity: 0.34
cat("Standard k-means purity:", round(purity_standard, 3), "\n")
#> Standard k-means purity: 0.8
cat("\n--- Elastic K-Means ---\n")
#>
#> --- Elastic K-Means ---
print(table(Elastic = ekm$labels, True = cls_sub))
#> True
#> Elastic aa ao dcl iy sh
#> 1 7 10 7 10 9
#> 2 0 0 1 0 0
#> 3 0 0 0 0 1
#> 4 0 0 2 0 0
#> 5 3 0 0 0 0
cat("\n--- Standard K-Means ---\n")
#>
#> --- Standard K-Means ---
print(table(Standard = km_std$cluster, True = cls_sub))
#> True
#> Standard aa ao dcl iy sh
#> 1 1 8 0 0 0
#> 2 0 0 10 0 0
#> 3 9 0 0 0 0
#> 4 0 0 0 7 4
#> 5 0 2 0 3 6
df_purity <- data.frame(
method = c("Standard K-Means", "Elastic K-Means"),
purity = c(purity_standard, purity_elastic)
)
ggplot(df_purity, aes(x = reorder(.data$method, .data$purity),
y = .data$purity, fill = .data$method)) +
geom_col(width = 0.5) +
geom_text(aes(label = sprintf("%.1f%%", .data$purity * 100)),
hjust = -0.2, size = 4) +
scale_fill_manual(values = c("Standard K-Means" = "#4A90D9",
"Elastic K-Means" = "#D55E00")) +
coord_flip(ylim = c(0, 1.1)) +
labs(title = "Cluster Purity: Elastic vs Standard",
x = "", y = "Purity") +
guides(fill = "none")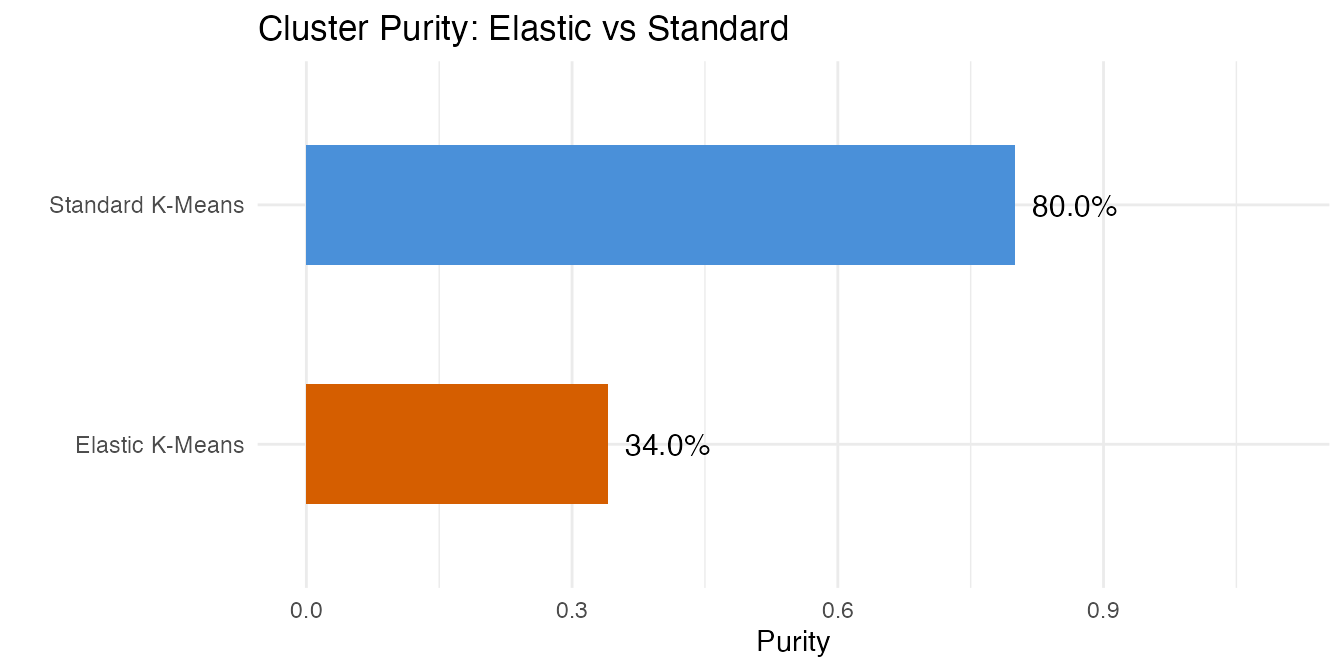
Standard k-means typically achieves higher purity on this spectral dataset. The elastic approach groups curves by shape similarity (modulo warping), but for spectral data the frequency positions of peaks carry the discriminative signal that warping removes.
9. Hierarchical View
A dendrogram reveals relationships between phoneme classes at multiple granularities.
ehc <- elastic.hclust(fd_sub, method = "complete")
print(ehc)
#> Elastic Hierarchical Clustering
#> Curves: 50 x 150 grid points
#> Method: complete
#> Merge steps: 49
plot(ehc) +
labs(title = "Elastic Hierarchical Clustering Dendrogram",
subtitle = "Complete linkage on Fisher-Rao distances")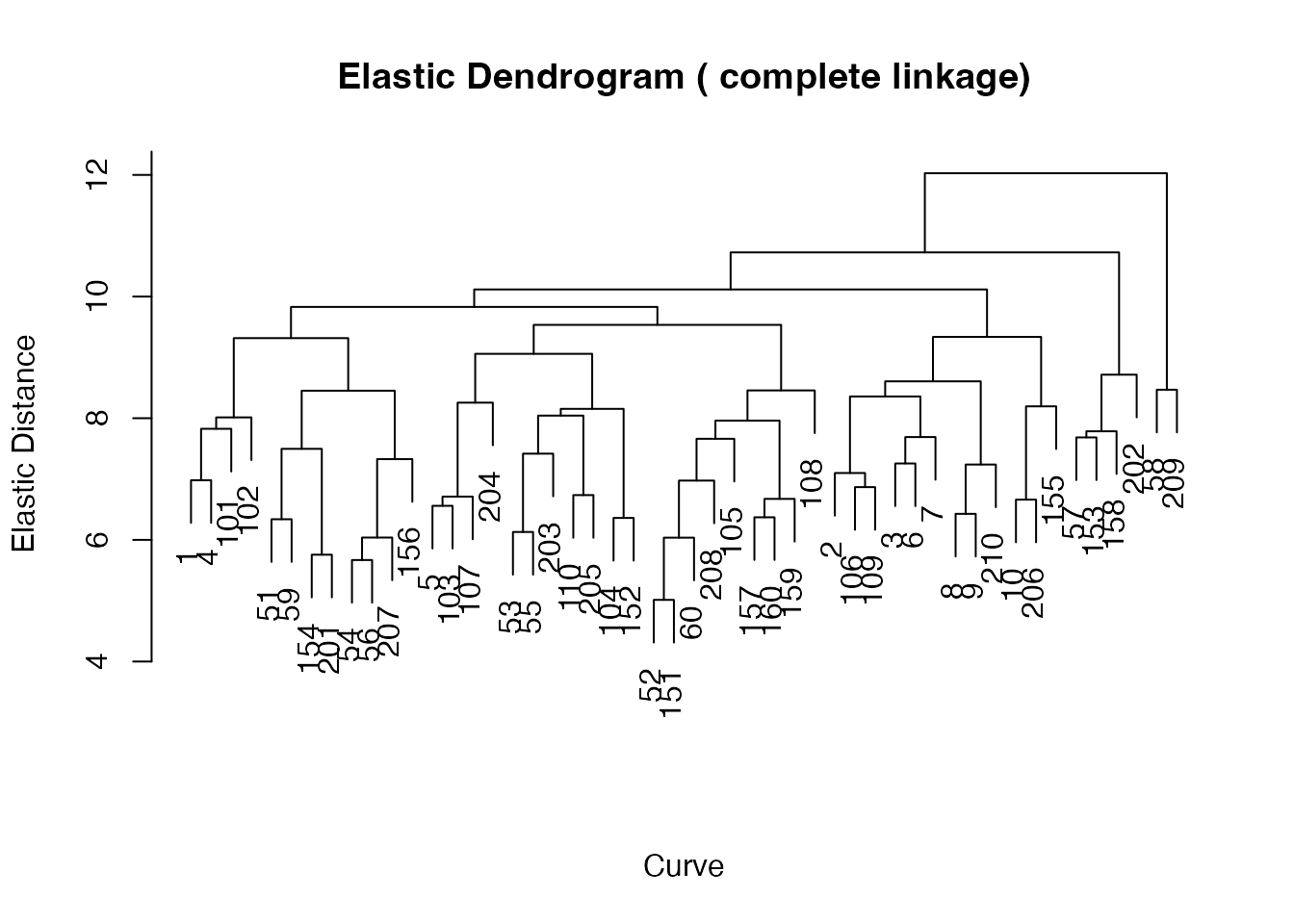
#> NULL
hc_labels <- elastic.cutree(ehc, k = 5)
purity_hc <- purity(hc_labels, cls_sub)
cat("Hierarchical clustering purity:", round(purity_hc, 3), "\n")
#> Hierarchical clustering purity: 0.4
cat("\n--- Hierarchical Clustering (k = 5) ---\n")
#>
#> --- Hierarchical Clustering (k = 5) ---
print(table(HClust = hc_labels, True = cls_sub))
#> True
#> HClust aa ao dcl iy sh
#> 1 2 4 2 2 2
#> 2 7 0 2 1 2
#> 3 1 4 6 5 4
#> 4 0 1 0 2 1
#> 5 0 1 0 0 1The dendrogram shows the hierarchical structure of spectral similarity. Cutting at k = 5 produces clusters whose purity is comparable to elastic k-means (both lower than standard k-means for this spectral dataset).
df_summary <- data.frame(
Method = c("Standard K-Means (L2)", "Elastic K-Means",
"Elastic Hierarchical"),
Purity = c(purity_standard, purity_elastic, purity_hc)
)
knitr::kable(df_summary, digits = 3,
caption = "Clustering Purity Comparison (50 curves, 5 classes)")| Method | Purity |
|---|---|
| Standard K-Means (L2) | 0.80 |
| Elastic K-Means | 0.34 |
| Elastic Hierarchical | 0.40 |
10. Conclusions
This example illustrates an important lesson: elastic methods are not universally superior. On phoneme log-periodograms, standard Euclidean k-means achieves higher clustering purity than elastic k-means or hierarchical clustering.
The reason is domain-specific. Elastic alignment via SRSF/Fisher-Rao warps the independent variable (here: frequency) to match curve shapes. For time-domain signals with phase variation (e.g., gait data, ECG), this is exactly what you want — it factors out irrelevant timing differences. But for spectral data, the frequency position of peaks IS the signal. A peak at 500 Hz vs 1000 Hz distinguishes vowels; warping the frequency axis destroys this information.
When to use elastic methods: - Time-domain curves with nuisance phase variation (gait, growth, ECG) - Data where shape similarity matters more than alignment
When standard metrics are better: - Spectral data (frequency-domain) where peak positions are discriminative - Data where the independent variable has physical meaning that should not be warped
The shape distance matrix and MDS embedding remain useful for visualization and understanding spectral structure, even when elastic clustering does not outperform standard methods.
See Also
-
vignette("articles/shape-analysis")— shape space theory, orbit representatives, and shape distance computation -
vignette("articles/elastic-clustering")— elastic k-means and hierarchical clustering on simulated data -
vignette("articles/distance-metrics")— comparison of functional distance measures
References
Ferraty, F. and Vieu, P. (2006). Nonparametric Functional Data Analysis. Springer. Chapter on the phoneme dataset.
Srivastava, A. and Klassen, E. (2016). Functional and Shape Data Analysis. Springer.
Kurtek, S., Srivastava, A., Klassen, E. and Ding, Z. (2012). Statistical Modeling of Curves Using Shapes and Related Features. Journal of the American Statistical Association, 107(499), 1152–1165.