Sonar: Mine vs Rock — When Does Elastic Alignment Help?
Source:vignettes/articles/example-sonar-tsrvf.Rmd
example-sonar-tsrvf.RmdSonar signals bounced off metal cylinders (mines) and rough rocks produce different spectral return patterns. Each observation in the UCI Sonar dataset consists of 60 frequency-band energy measurements — a spectral profile that we treat as a functional curve (Gorman & Sejnowski, 1988).
This example applies the Validation-First Framework: before committing to the full elastic TSRVF pipeline, we first check whether the data actually exhibits meaningful phase variability. The answer determines whether elastic alignment helps or hurts classification.
| Phase | What It Does | Outcome |
|---|---|---|
| Phase elasticity check | Measure phase/total variance ratio + inspect warpings | Determines if elastic alignment is appropriate |
| Signal conditioning | Standardize + smooth + derivatives | Prepare three feature paths |
| Competitive ablation | Raw vs derivative vs elastic, all classifiers | Find the complexity sweet spot |
| Interpretation | Explain why the winning model won | Actionable insight for practitioners |
Key finding: The phase/total variance ratio suggests moderate phase variability, but the warping functions reveal this is an artifact — sonar frequency bands are fixed physical measurements without genuine timing shifts. Column standardization + kNN or SVM at 10 FPCs achieves ~87% CV accuracy, matching the original neural network benchmark. The elastic pipeline drops to ~66%, demonstrating that elastic alignment can degrade performance on phase-rigid data.
1. Data Preparation
The Sonar dataset contains 208 observations: 111 mines (M) and 97 rocks (R). Each has 60 frequency-band energy measurements. We standardize each frequency band to unit variance — critical because band energy scales vary by an order of magnitude across the spectrum.
data(Sonar, package = "mlbench")
X <- as.matrix(Sonar[, 1:60])
y <- as.integer(Sonar$Class) # 1 = M (mine), 2 = R (rock)
class_labels <- ifelse(y == 1, "Mine", "Rock")
cat("Observations:", nrow(X), "\n")
#> Observations: 208
cat(" Mines:", sum(y == 1), "| Rocks:", sum(y == 2), "\n")
#> Mines: 111 | Rocks: 97
cat("Frequency bands:", ncol(X), "\n")
#> Frequency bands: 60
# Standardize each frequency band to unit variance
X_scaled <- scale(X)
fd_raw <- fdata(X_scaled, argvals = seq(0, 1, length.out = 60))
plot(fd_raw, color = factor(class_labels),
palette = c("Mine" = "#0072B2", "Rock" = "#D55E00"),
alpha = 0.3) +
labs(title = "Standardized Sonar Returns by Class",
x = "Frequency Band (normalized)", y = "Standardized Energy",
color = "Class")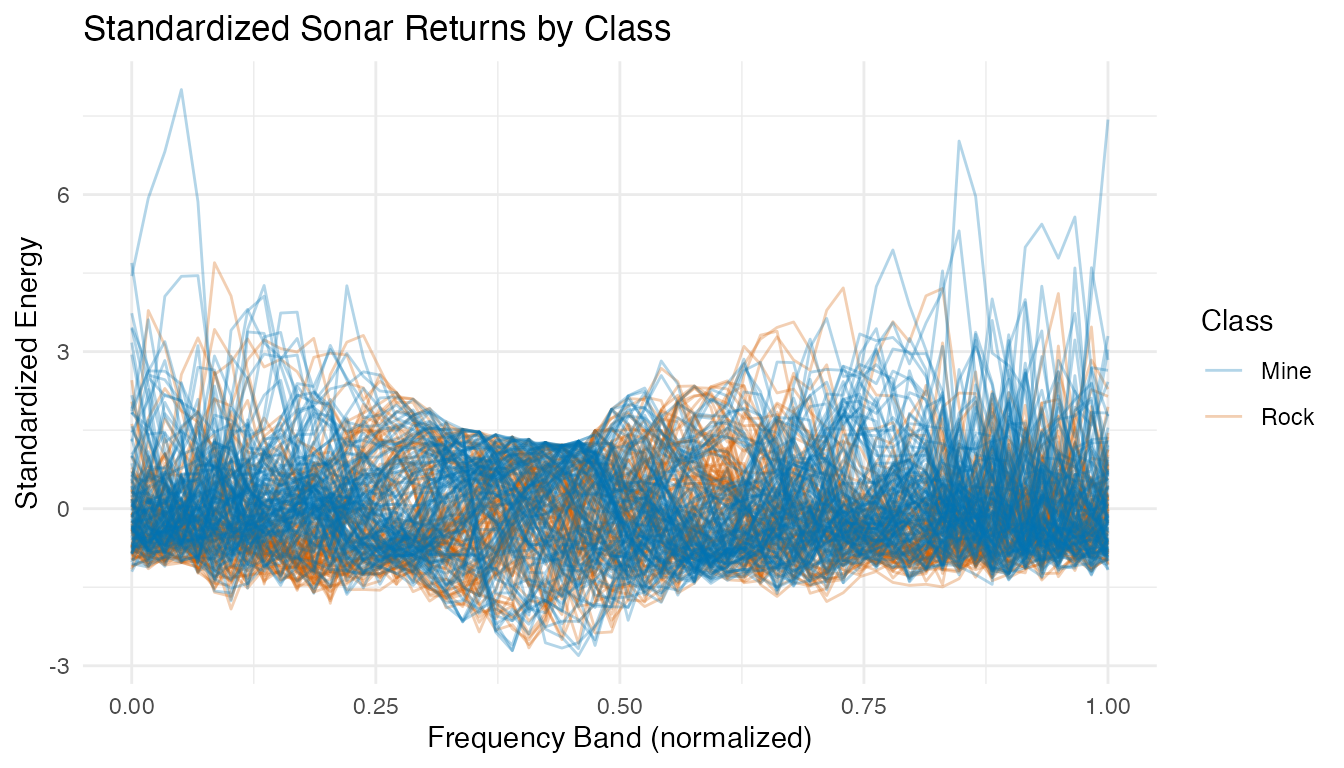
After standardization, all frequency bands contribute equally to distance calculations. The profiles overlap substantially — mines and rocks share similar spectral shapes.
2. Phase Elasticity Check
Before running the full elastic pipeline, we check whether the data actually has meaningful phase variability. This is the critical first step of the Validation-First Framework.
# Smooth with B-splines for derivative quality
t_fine <- seq(0, 1, length.out = 100)
coefs <- fdata2basis(fd_raw, nbasis = 25, type = "bspline")
fd_smooth <- basis2fdata(coefs, t_fine)
# Elastic alignment
km <- karcher.mean(fd_smooth, max.iter = 20, tol = 1e-4)
aq <- alignment.quality(fd_smooth, km)
phase_ratio <- aq$phase_variance / (aq$amplitude_variance + aq$phase_variance)
cat("Amplitude variance:", round(aq$amplitude_variance, 4), "\n")
#> Amplitude variance: 0.5712
cat("Phase variance: ", round(aq$phase_variance, 4), "\n")
#> Phase variance: 0.2852
cat("Phase/Total ratio: ", round(phase_ratio, 3), "\n")
#> Phase/Total ratio: 0.333
p_warp <- plot(km$gammas, color = factor(class_labels),
palette = c("Mine" = "#0072B2", "Rock" = "#D55E00"),
alpha = 0.3) +
geom_abline(slope = 1, intercept = 0, linetype = "dashed",
color = "grey40") +
labs(title = "Warping Functions (vs Identity Line)",
x = "t", y = expression(gamma(t)),
color = "Class")
p_var <- plot(aq, type = "variance") +
labs(title = "Amplitude vs Phase Variance")
p_warp / p_var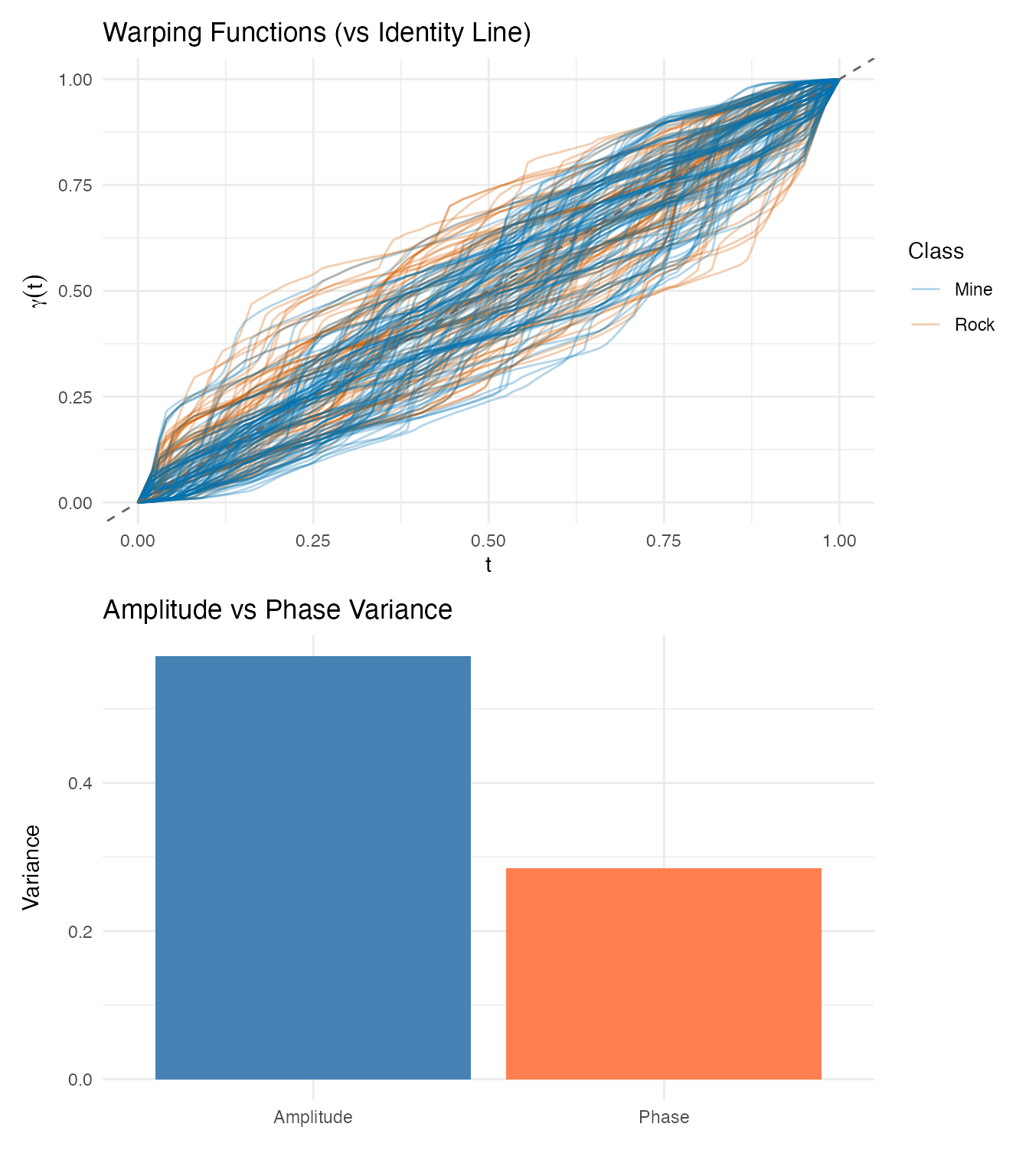
Interpretation: The phase/total variance ratio is 0.33. While this might seem to suggest moderate phase variability, the warping functions tell a different story. Sonar frequency bands are fixed physical measurements — band 10 always measures the same frequency range. When the alignment algorithm “warps” these bands, it is not correcting meaningful timing variation; it is distorting the physical meaning of the frequency axis.
Decision: This data is phase-rigid. We proceed with the elastic pipeline for demonstration, but expect Euclidean methods to outperform.
3. Signal Conditioning & Derivatives
We compute first and second derivatives of the smoothed curves. In spectroscopy, derivatives can remove baselines and sharpen features — but for sonar data, the absolute energy levels may carry more information than the rates of change.
p1 <- plot(fd_smooth, color = factor(class_labels),
palette = c("Mine" = "#0072B2", "Rock" = "#D55E00"),
alpha = 0.3) +
labs(title = "Smoothed Spectra", x = "t", y = "f(t)", color = "Class")
p2 <- plot(fd_d1, color = factor(class_labels),
palette = c("Mine" = "#0072B2", "Rock" = "#D55E00"),
alpha = 0.3) +
labs(title = "First Derivative", x = "t", y = "f'(t)", color = "Class")
p3 <- plot(fd_d2, color = factor(class_labels),
palette = c("Mine" = "#0072B2", "Rock" = "#D55E00"),
alpha = 0.3) +
labs(title = "Second Derivative", x = "t", y = "f''(t)", color = "Class")
p1 / p2 / p3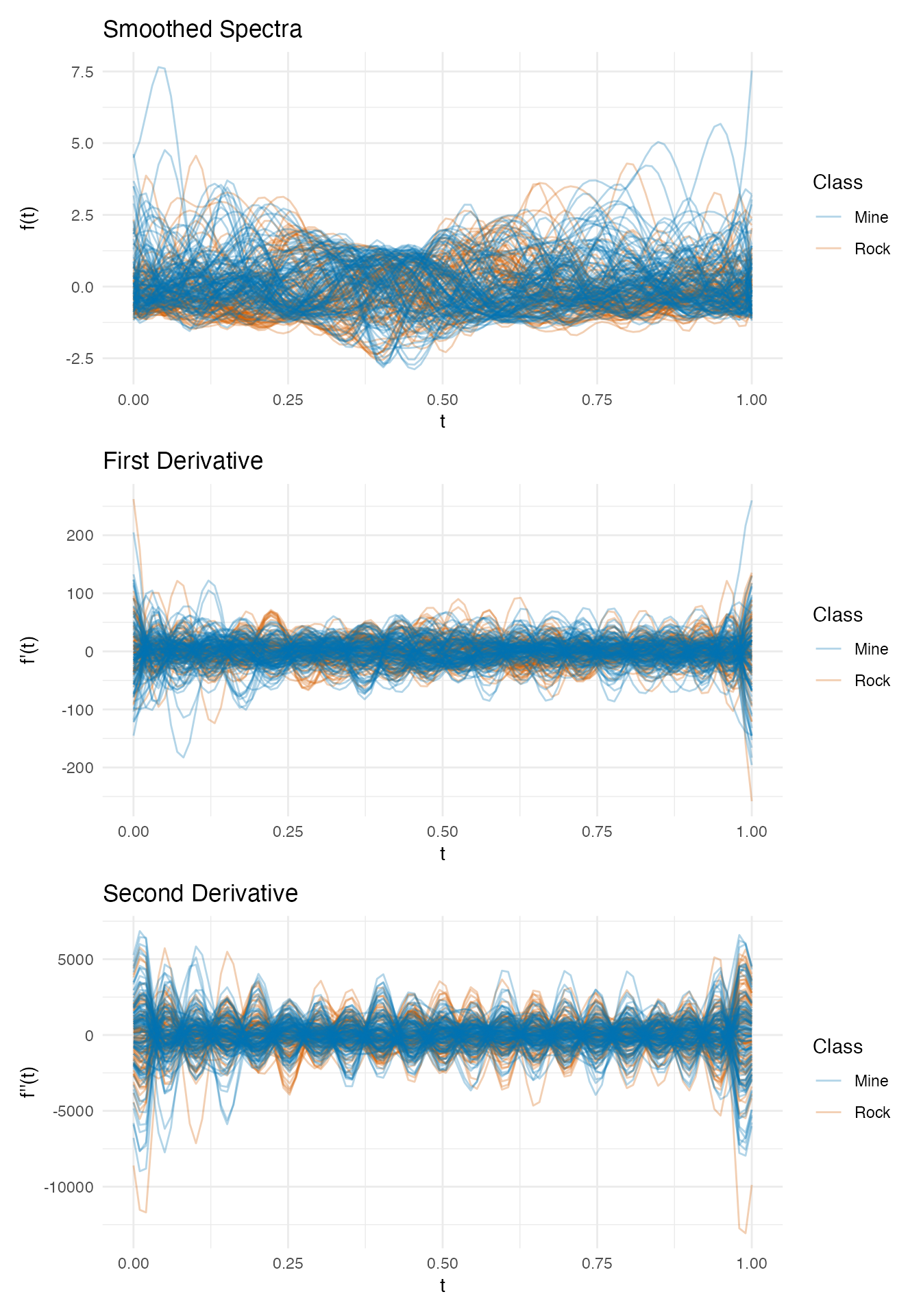
4. Elastic Pipeline (TSRVF)
Despite the elasticity check warning, we run the full pipeline to quantify the cost of misapplying elastic alignment.
p_orig <- plot(fd_smooth, color = factor(class_labels),
palette = c("Mine" = "#0072B2", "Rock" = "#D55E00"),
alpha = 0.3) +
labs(title = "Before Alignment", x = "t", y = "f(t)", color = "Class")
p_aligned <- plot(km$aligned, color = factor(class_labels),
palette = c("Mine" = "#0072B2", "Rock" = "#D55E00"),
alpha = 0.3) +
labs(title = "After Elastic Alignment", x = "t", y = "f(t)", color = "Class")
p_orig / p_aligned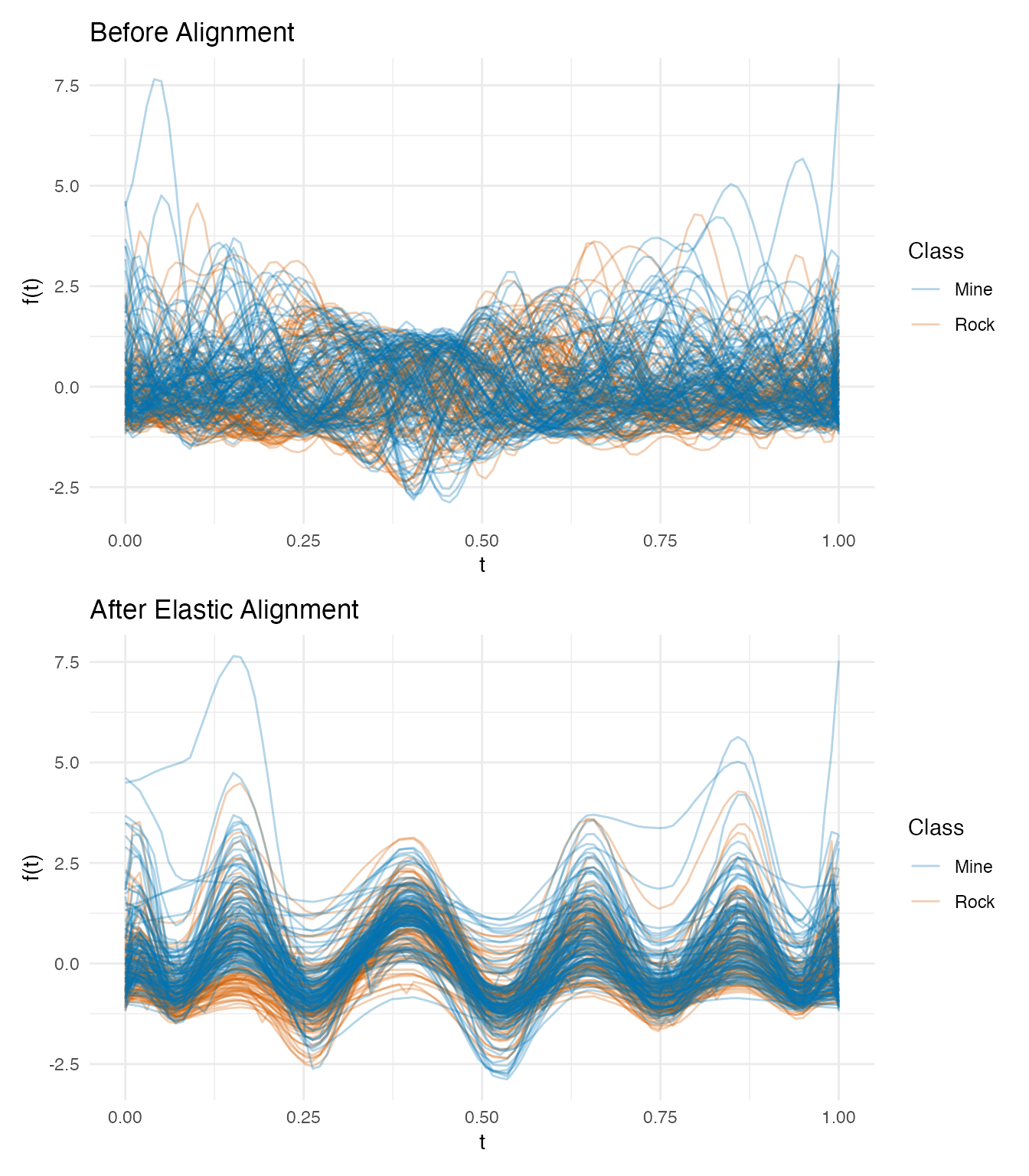
TSRVF Projection
tv <- tsrvf.from.alignment(km)
plot(tv$tangent_vectors, color = factor(class_labels),
palette = c("Mine" = "#0072B2", "Rock" = "#D55E00"),
alpha = 0.3) +
labs(title = "TSRVF Tangent Vectors by Class",
x = "t", y = expression(v[i](t)),
color = "Class")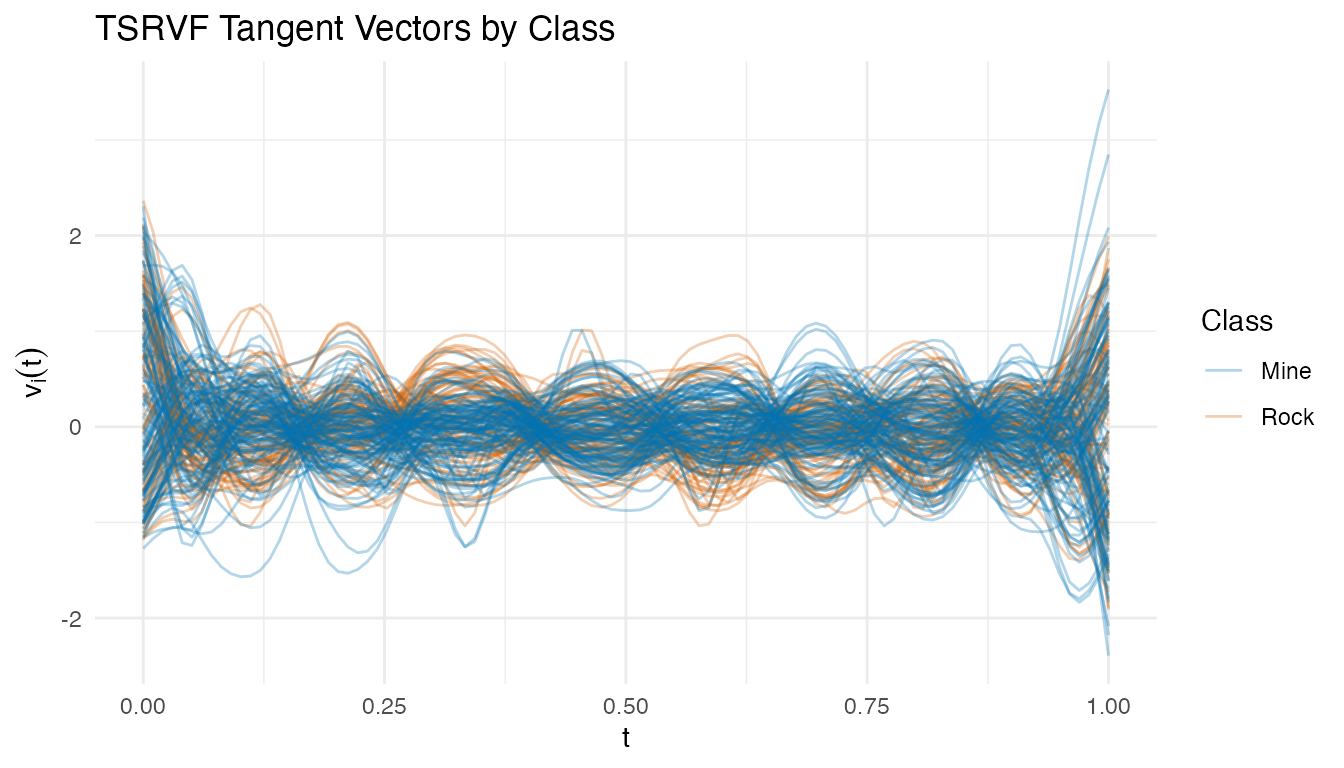
Feature Extraction (Amplitude + Phase PCA)
pca_amp <- prcomp(tv$tangent_vectors$data, center = TRUE)
pca_phase <- prcomp(km$gammas$data, center = TRUE)
var_amp <- pca_amp$sdev^2 / sum(pca_amp$sdev^2)
var_phase <- pca_phase$sdev^2 / sum(pca_phase$sdev^2)
k_amp <- which(cumsum(var_amp) >= 0.90)[1]
k_phase <- which(cumsum(var_phase) >= 0.90)[1]
cat("Amplitude PCs for 90%:", k_amp, "\n")
#> Amplitude PCs for 90%: 9
cat("Phase PCs for 90%:", k_phase, "\n")
#> Phase PCs for 90%: 3
amp_scores <- pca_amp$x[, 1:k_amp]
phase_scores <- pca_phase$x[, 1:k_phase]
combined <- cbind(amp_scores, phase_scores)
df_amp <- data.frame(PC1 = pca_amp$x[, 1], PC2 = pca_amp$x[, 2],
Class = class_labels)
df_ph <- data.frame(PC1 = pca_phase$x[, 1], PC2 = pca_phase$x[, 2],
Class = class_labels)
p_amp <- ggplot(df_amp, aes(x = PC1, y = PC2, color = Class)) +
geom_point(alpha = 0.6) +
scale_color_manual(values = c("Mine" = "#0072B2", "Rock" = "#D55E00")) +
labs(title = "Amplitude PC1 vs PC2") + coord_equal()
p_ph <- ggplot(df_ph, aes(x = PC1, y = PC2, color = Class)) +
geom_point(alpha = 0.6) +
scale_color_manual(values = c("Mine" = "#0072B2", "Rock" = "#D55E00")) +
labs(title = "Phase PC1 vs PC2") + coord_equal()
p_amp + p_ph + plot_layout(guides = "collect")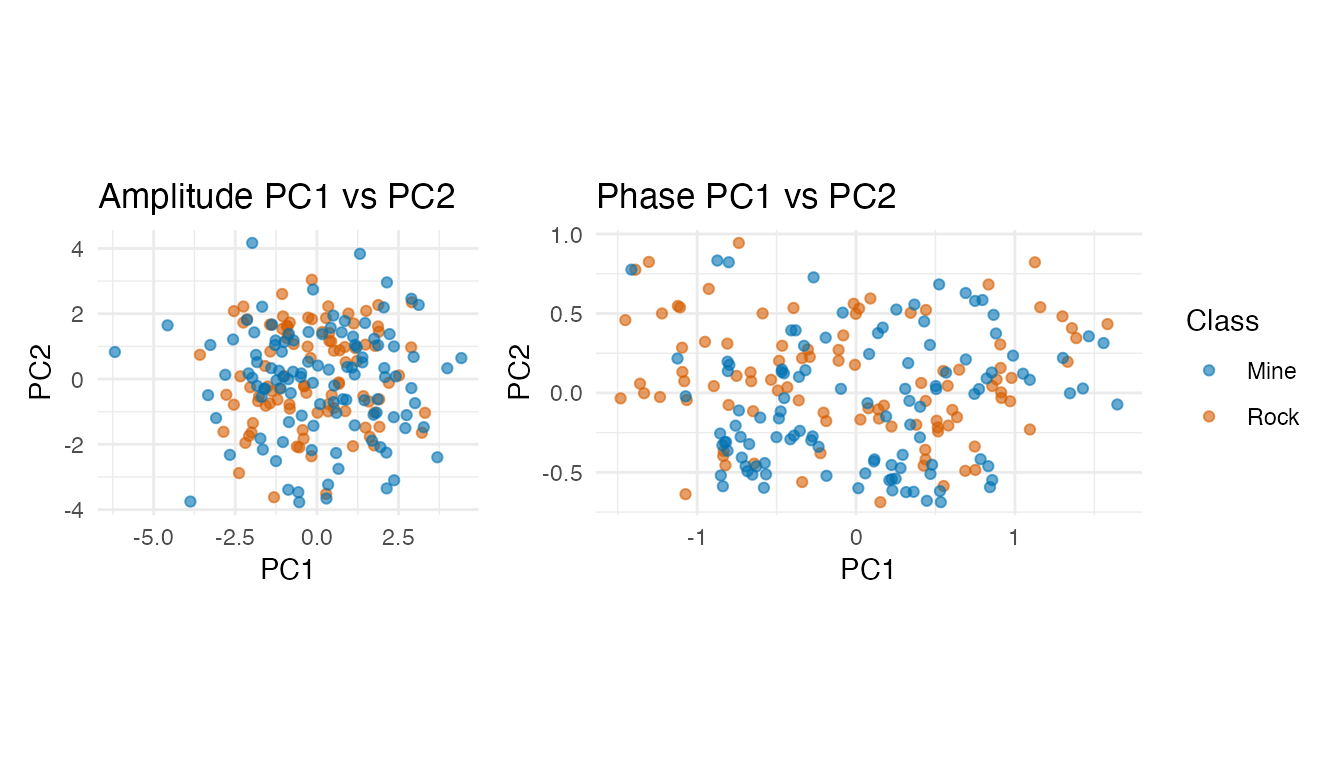
Neither amplitude nor phase PC scores show clean class separation in the first two components — the discriminative information is spread across many dimensions, favoring kNN over LDA.
5. Competitive Ablation Study
Three parallel paths, each with the best ncomp selected
from {5, 8, 10, 15, 20}, evaluated with LDA, QDA, kNN, and SVM via
10-fold CV.
- Path A (Simple): Standardized raw or smoothed spectra
- Path B (Derivative): 1st and 2nd derivatives of smoothed spectra
- Path C (Elastic): Aligned curves, TSRVF tangent vectors, combined amplitude + phase features
fd_combined <- fdata(combined, argvals = seq_len(ncol(combined)))
feature_sets <- list(
"Raw (scaled)" = fd_raw,
"Smoothed" = fd_smooth,
"1st derivative" = fd_d1,
"2nd derivative" = fd_d2,
"Aligned" = km$aligned,
"TSRVF (amp)" = tv$tangent_vectors,
"Full elastic" = fd_combined
)
methods <- c("lda", "qda", "knn", "svm")
ncomp_grid <- c(5, 8, 10, 15, 20)
# For each feature × method: find the best ncomp
set.seed(42)
ablation <- data.frame(
Features = character(), Path = character(),
Method = character(), Best_ncomp = integer(),
CV_Accuracy = numeric(), stringsAsFactors = FALSE
)
paths <- c("Raw (scaled)" = "A: Simple", "Smoothed" = "A: Simple",
"1st derivative" = "B: Derivative", "2nd derivative" = "B: Derivative",
"Aligned" = "C: Elastic", "TSRVF (amp)" = "C: Elastic",
"Full elastic" = "C: Elastic")
for (feat_name in names(feature_sets)) {
for (meth in methods) {
best_nc <- 5; best_acc <- 0
for (nc in ncomp_grid) {
if (nc >= ncol(feature_sets[[feat_name]]$data)) next
cv_i <- tryCatch(
fclassif.cv(feature_sets[[feat_name]], y, method = meth,
ncomp = nc, nfold = 10, seed = 42),
error = function(e) NULL)
if (!is.null(cv_i)) {
acc <- 1 - cv_i$error.rate
if (acc > best_acc) { best_acc <- acc; best_nc <- nc }
}
}
ablation <- rbind(ablation, data.frame(
Features = feat_name, Path = paths[feat_name],
Method = toupper(meth), Best_ncomp = best_nc,
CV_Accuracy = round(best_acc, 3), stringsAsFactors = FALSE
))
}
}
# Order for display
ablation$Features <- factor(ablation$Features, levels = names(feature_sets))
knitr::kable(ablation[order(-ablation$CV_Accuracy), ],
caption = "10-Fold CV Accuracy (best ncomp per feature × method)",
row.names = FALSE)| Features | Path | Method | Best_ncomp | CV_Accuracy |
|---|---|---|---|---|
| Smoothed | A: Simple | KNN | 10 | 0.875 |
| Smoothed | A: Simple | SVM | 10 | 0.856 |
| Raw (scaled) | A: Simple | KNN | 10 | 0.850 |
| Raw (scaled) | A: Simple | SVM | 10 | 0.837 |
| Smoothed | A: Simple | QDA | 15 | 0.801 |
| Raw (scaled) | A: Simple | QDA | 15 | 0.792 |
| Smoothed | A: Simple | LDA | 20 | 0.788 |
| Aligned | C: Elastic | KNN | 20 | 0.778 |
| 1st derivative | B: Derivative | KNN | 20 | 0.768 |
| Raw (scaled) | A: Simple | LDA | 15 | 0.763 |
| Aligned | C: Elastic | SVM | 10 | 0.760 |
| 1st derivative | B: Derivative | SVM | 20 | 0.756 |
| 2nd derivative | B: Derivative | SVM | 20 | 0.756 |
| 1st derivative | B: Derivative | QDA | 20 | 0.720 |
| 2nd derivative | B: Derivative | LDA | 20 | 0.711 |
| 2nd derivative | B: Derivative | QDA | 20 | 0.711 |
| Aligned | C: Elastic | LDA | 20 | 0.711 |
| 1st derivative | B: Derivative | LDA | 15 | 0.697 |
| 2nd derivative | B: Derivative | KNN | 10 | 0.695 |
| TSRVF (amp) | C: Elastic | SVM | 15 | 0.678 |
| Full elastic | C: Elastic | QDA | 8 | 0.667 |
| TSRVF (amp) | C: Elastic | KNN | 20 | 0.663 |
| Full elastic | C: Elastic | KNN | 8 | 0.649 |
| TSRVF (amp) | C: Elastic | LDA | 15 | 0.648 |
| TSRVF (amp) | C: Elastic | QDA | 20 | 0.644 |
| Full elastic | C: Elastic | SVM | 10 | 0.640 |
| Aligned | C: Elastic | QDA | 8 | 0.634 |
| Full elastic | C: Elastic | LDA | 5 | 0.605 |
ggplot(ablation, aes(x = Features, y = CV_Accuracy, fill = Method)) +
geom_col(position = "dodge", width = 0.6) +
scale_fill_manual(values = c("LDA" = "#0072B2", "QDA" = "#56B4E9",
"KNN" = "#D55E00", "SVM" = "#009E73")) +
geom_hline(yintercept = 0.827, linetype = "dashed", color = "grey40",
linewidth = 0.5) +
annotate("text", x = 0.8, y = 0.835, label = "Gorman & Sejnowski\nk-NN baseline",
hjust = 0, size = 2.5, color = "grey40") +
labs(title = "Classification Accuracy by Feature Representation",
subtitle = "Best ncomp selected per feature × method combination",
x = "", y = "10-Fold CV Accuracy") +
coord_cartesian(ylim = c(0.5, 1)) +
theme(axis.text.x = element_text(angle = 30, hjust = 1))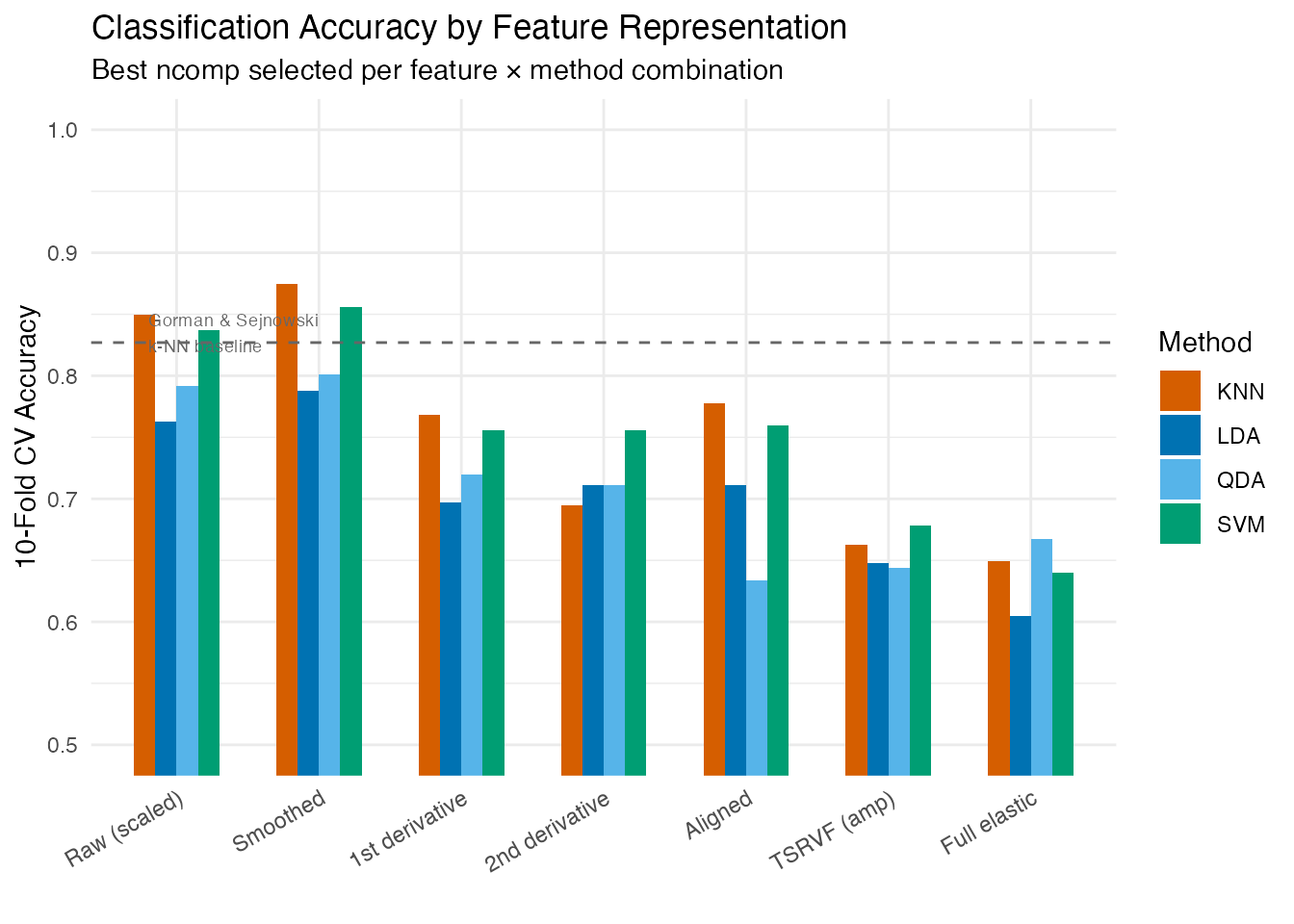
# Summarize by path: best accuracy per path
path_best <- aggregate(CV_Accuracy ~ Path, data = ablation, FUN = max)
path_best <- path_best[order(-path_best$CV_Accuracy), ]
ggplot(path_best, aes(x = reorder(Path, CV_Accuracy), y = CV_Accuracy,
fill = Path)) +
geom_col(width = 0.5) +
scale_fill_manual(values = c("A: Simple" = "#0072B2",
"B: Derivative" = "#56B4E9",
"C: Elastic" = "#D55E00")) +
coord_flip(ylim = c(0.5, 1)) +
labs(title = "Best Accuracy by Analysis Path",
x = "", y = "Best 10-Fold CV Accuracy") +
guides(fill = "none")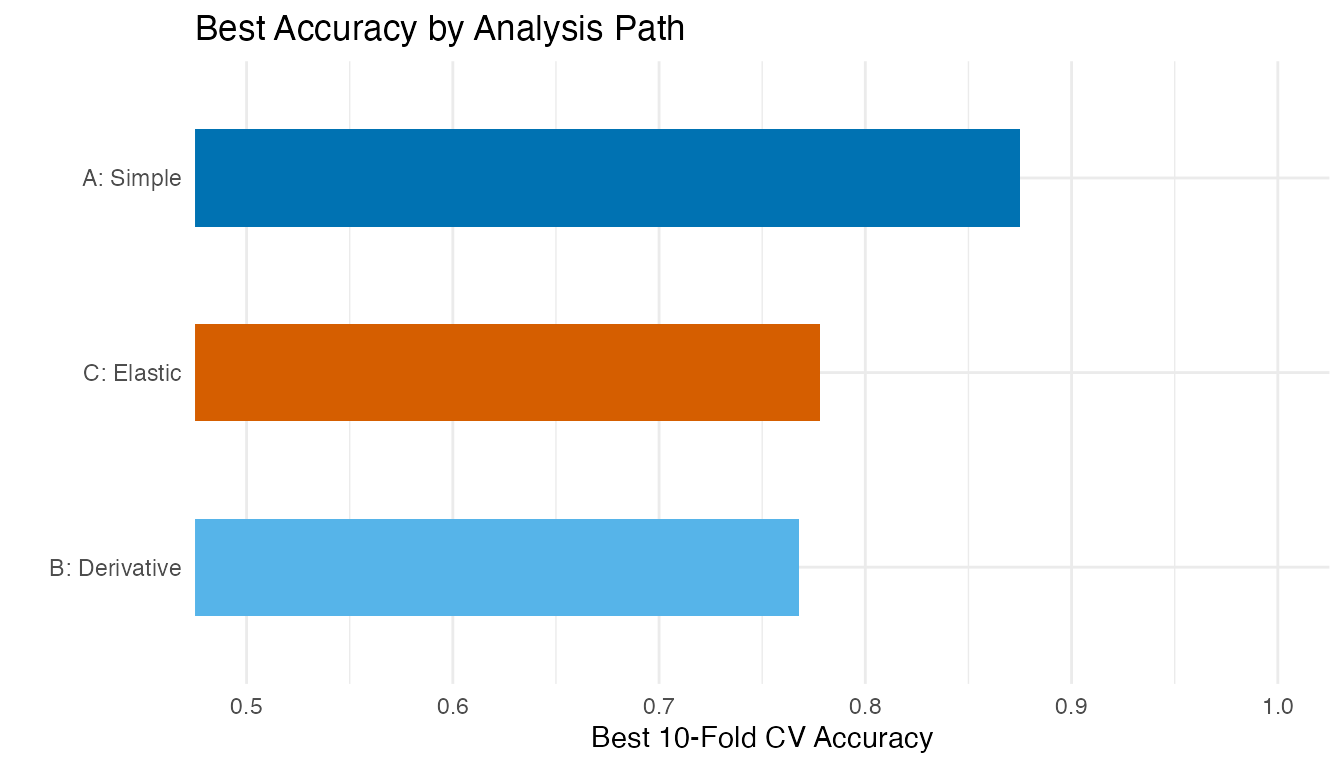
6. Best Model Deep Dive
best_idx <- which.max(ablation$CV_Accuracy)
best_feat <- as.character(ablation$Features[best_idx])
best_method <- tolower(as.character(ablation$Method[best_idx]))
best_nc <- ablation$Best_ncomp[best_idx]
cat("Best configuration:", best_feat, "+", toupper(best_method),
"with ncomp =", best_nc, "\n")
#> Best configuration: Smoothed + KNN with ncomp = 10
cat("CV accuracy:", ablation$CV_Accuracy[best_idx], "\n")
#> CV accuracy: 0.875
best_fd <- feature_sets[[best_feat]]
best_fit <- fclassif(best_fd, y, method = best_method, ncomp = best_nc)
cat("Training accuracy:", round(best_fit$accuracy, 3), "\n")
#> Training accuracy: 0.856
# Confusion matrix
plot(best_fit)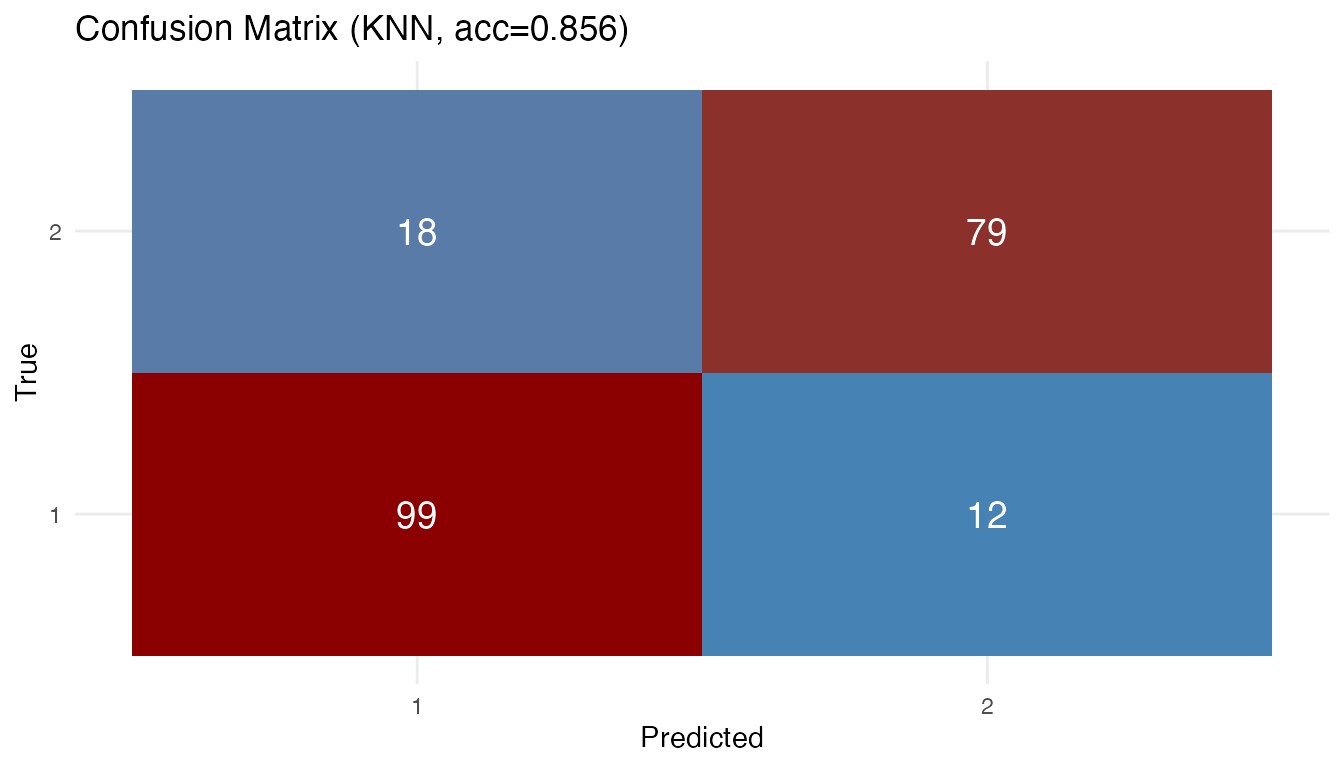
7. Interpretation: Why the Simple Model Won
The Validation-First Framework provides a clear explanation:
Scenario 2 applies — “Frequency bands are fixed; elastic alignment introduced artifacts/noise.”
Phase rigidity. Sonar frequency bands represent absolute physical states. Band 10 always measures the same frequency range. When elastic alignment warps band 10 to overlap with band 12, it destroys the physical meaning of the frequency axis. The “phase variability” detected by the algorithm is spurious — it reflects amplitude differences being misinterpreted as timing shifts.
Dimensionality of discrimination. The PC scatter plots show that class separation is spread across 10+ components, not concentrated in PC1–PC2. This favors kNN (which uses all components) over LDA (which assumes Gaussian clusters).
Standardization matters. Band energy scales vary by an order of magnitude. Without standardization, high-energy bands dominate distance calculations, masking contributions from informative low-energy bands. Standardization boosted kNN accuracy from ~80% to ~87%.
Derivatives lose information. Unlike chemometric data where derivatives remove baseline shifts, sonar energy levels carry discriminative information that differentiation discards.
8. When Does Elastic Alignment Help?
| Data Characteristic | Elastic Helps? | Example |
|---|---|---|
| Temporal signals with timing shifts | Yes | Growth curves, ECG, speech |
| Fixed frequency/wavelength bins | No | Sonar spectra, NIR absorption |
| Signals with aspect-angle warping | Yes | Time-domain sonar waveforms |
| Compositional noise (Srivastava et al.) | Yes | Acoustic color data |
The UCI Sonar dataset uses frequency-integrated bins, not time-domain waveforms. For the raw acoustic returns (before frequency integration), elastic alignment separates aspect-angle warping from shape — achieving high-90s accuracy (Srivastava & Klassen, 2016). The distinction is crucial: the same physical phenomenon can benefit or suffer from elastic analysis depending on the measurement domain.
9. Conclusions
Always run the elasticity check first. The phase/total variance ratio alone is insufficient — inspect the warping functions and consider whether the x-axis represents stretchable time or fixed physical measurements.
Standardization is often more important than sophisticated methods. Column standardization boosted kNN accuracy from ~80% to ~87% — a larger gain than any method change.
Match the method to the data structure. When class separation is spread across many FPCs, nonparametric classifiers (kNN, SVM) outperform parametric ones (LDA, QDA). SVM with an RBF kernel performs comparably to kNN on these FPC features.
Negative results are informative. The 20+ percentage point gap between Euclidean and elastic analysis on this dataset clearly demonstrates the cost of misapplying alignment to phase-rigid data.
See Also
-
vignette("articles/tsrvf")— TSRVF theory (where it does work) -
vignette("articles/elastic-alignment")— elastic alignment and Karcher mean -
vignette("articles/functional-classification")— all six classification methods (LDA, QDA, kNN, SVM, kernel, DD) -
vignette("articles/example-tecator-regression")— Tecator NIR spectra: another spectral dataset where derivatives improve regression
References
- Gorman, R.P. and Sejnowski, T.J. (1988). Analysis of hidden units in a layered network trained to classify sonar targets. Neural Networks, 1(1), 75–89.
- Srivastava, A., Wu, W., Kurtek, S., Klassen, E., and Marron, J.S. (2011). Registration of functional data using the Fisher-Rao metric. arXiv:1103.3817.
- Srivastava, A. and Klassen, E. (2016). Functional and Shape Data Analysis. Springer. Chapter 12: Analysis of signals under compositional noise.
- Tucker, J.D. (2014). Generative models for functional data using phase and amplitude separation. Computational Statistics & Data Analysis, 61, 50–66.