Introduction
Standard scalar-on-function regression assumes curves are well-aligned before analysis. When functional predictors contain phase variation — differences in the timing of features like peaks, valleys, or inflection points — standard methods confound shape with timing. The estimated coefficient function becomes blurred, and predictive power drops.
scalar.on.shape() solves this by building a regression
model that is invariant to phase variation. It uses the
Fisher-Rao elastic framework to separate shape (amplitude after
alignment) from timing (warping), then regresses the scalar
response on the shape of each curve.
When to Use Scalar-on-Shape
| Scenario | Recommended method |
|---|---|
| Curves well-aligned, linear relationship | fregre.lm() |
| Curves misaligned, response depends on amplitude | elastic.regression() |
| Curves misaligned, response depends on shape (phase-invariant) | scalar.on.shape() |
| Unknown alignment quality | Compare all three |
Simulating Data with Phase Variation
We simulate a biomechanics-inspired scenario: 200 force–time curves where each curve is a single bump (e.g., a ground reaction force impulse during a vertical jump). The bump’s location varies across subjects (phase variation from different jump timings), but the scalar response — vertical jump height — depends on the bump’s width (a shape feature that is invariant to when the jump occurred).
This creates a challenging problem for standard FPCA-based regression: the first several principal components capture the dominant phase variation (where the bump is), not the shape variation (how wide it is). The shape signal is buried in higher-order components that are hard to select in practice.
set.seed(42)
n <- 200
m <- 60
t_grid <- seq(0, 1, length.out = m)
X <- matrix(0, n, m)
y <- numeric(n)
for (i in 1:n) {
# Shape: bump width (this predicts jump height)
width_param <- rnorm(1, 0, 1)
width <- 0.06 + 0.015 * width_param
# Phase: bump center varies continuously (dominant variation)
center <- runif(1, 0.25, 0.75)
X[i, ] <- exp(-(t_grid - center)^2 / (2 * width^2)) +
rnorm(m, sd = 0.015)
# Response depends only on width (shape), not center (phase)
y[i] <- 3 * width_param + rnorm(1, sd = 0.3)
}
fd <- fdata(X, argvals = t_grid)
plot(fd, main = "Simulated Ground Reaction Force Curves",
xlab = "Normalized Time", ylab = "Force (BW)")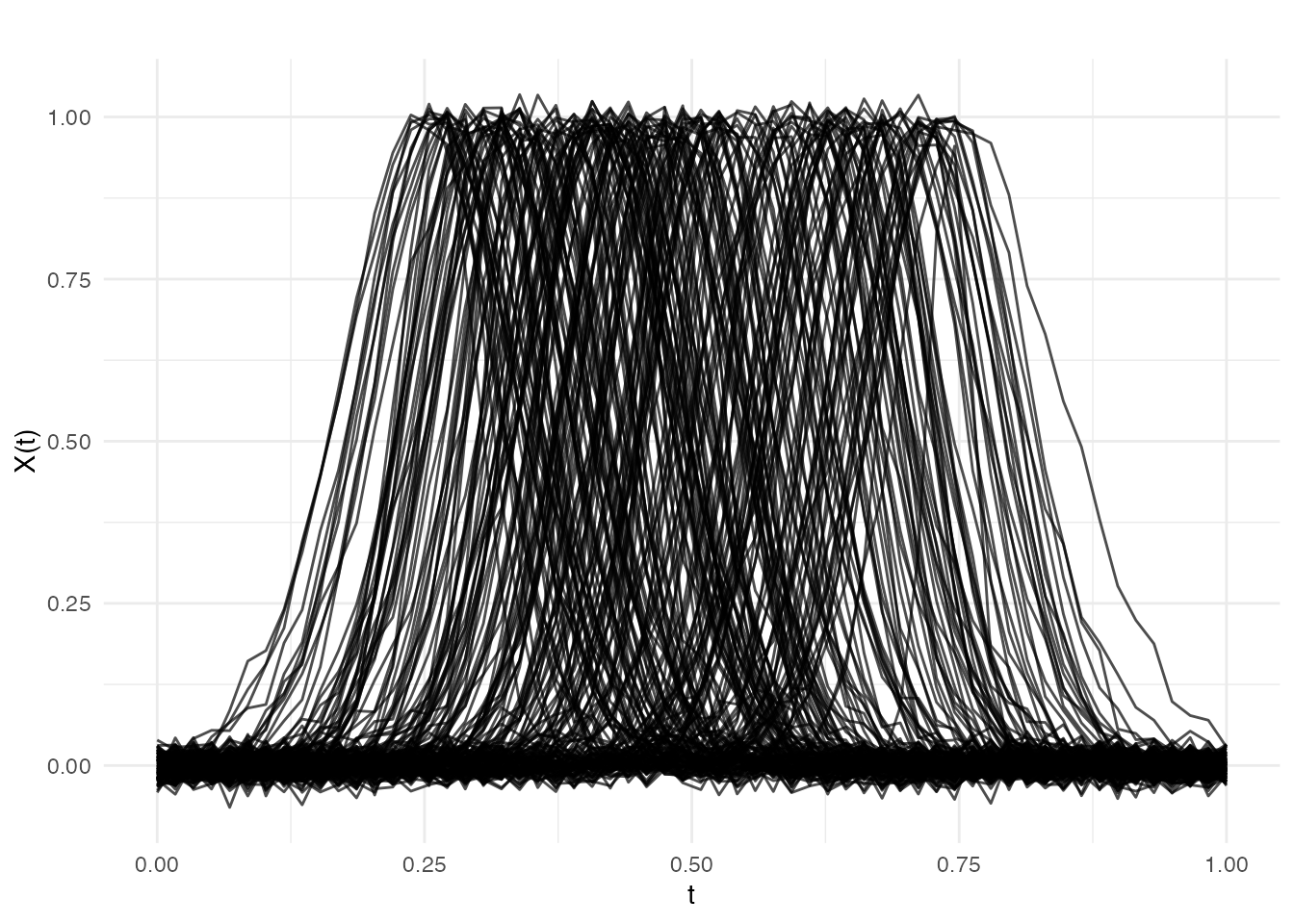
Note how the bumps are located at different positions along the time axis — this is phase variation. The response depends only on bump width, a shape feature that standard FPCA mixes with phase.
Fitting the Model
Identity Index (Linear Link)
The simplest model uses index.method = "identity", which
assumes a linear relationship between the shape score and the
response.
fit_id <- scalar.on.shape(fd, y, nbasis = 11, lambda = 1e-3,
index.method = "identity",
max.iter.outer = 15, tol = 1e-4)
print(fit_id)
#> Scalar-on-Shape Regression
#> Index method: identity
#> R-squared: 0.859
#> SSE: 228.8896
#> Iterations: 15 (outer), 15 (inner)Examining the Fitted Model
# Shape index function beta(t)
df_beta <- data.frame(t = t_grid, beta = fit_id$beta)
ggplot(df_beta, aes(x = t, y = beta)) +
geom_line(color = "#4A90D9", linewidth = 1.2) +
labs(title = "Estimated Shape Index Function",
subtitle = "beta(t): the shape pattern that predicts y",
x = "Normalized Time", y = expression(beta(t)))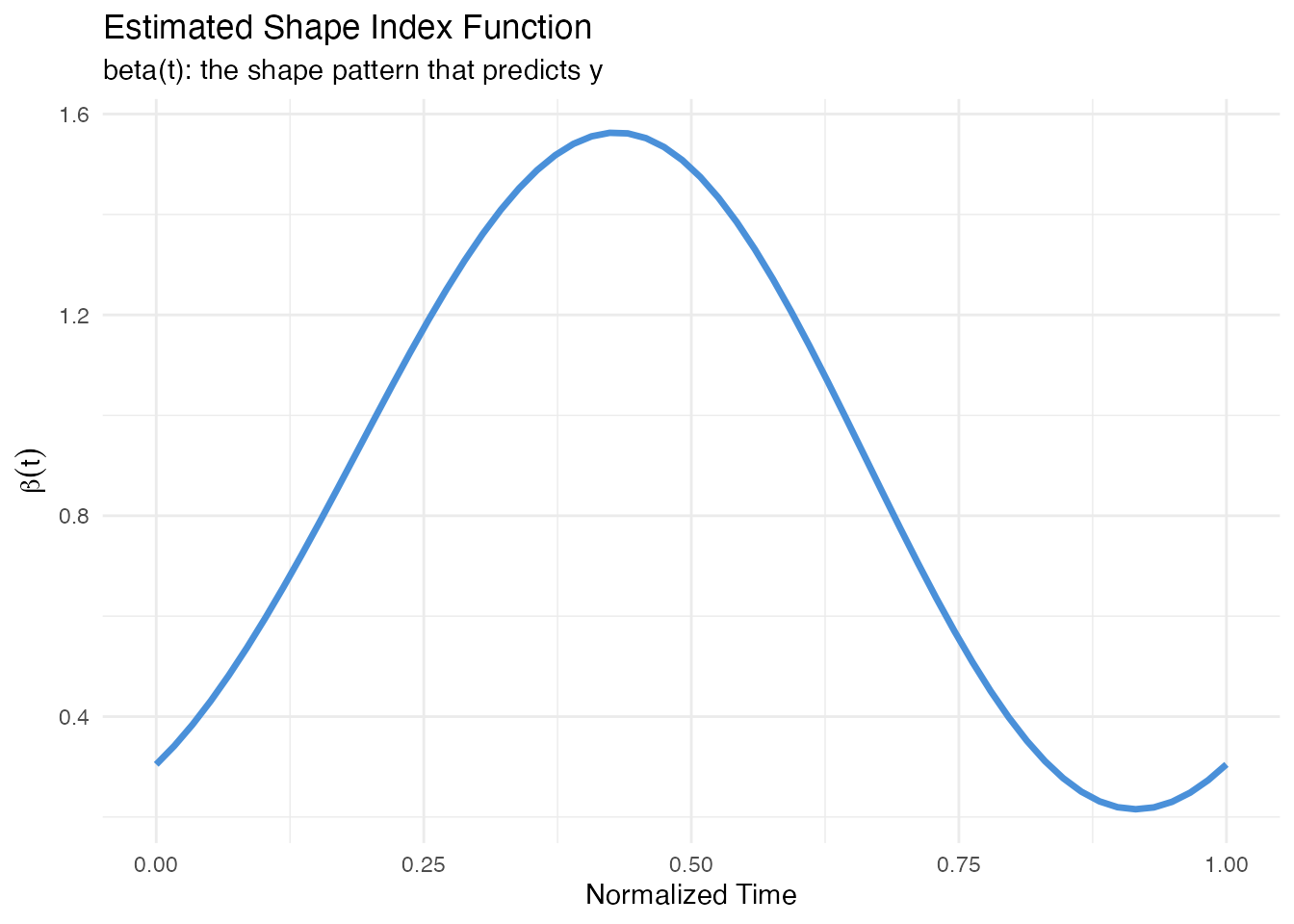
The shape index reveals which aspects of curve shape drive the response. Peaks in correspond to regions where the curve shape is most predictive.
# Shape scores vs observed response
df_scores <- data.frame(score = fit_id$shape.scores, y = y)
ggplot(df_scores, aes(x = score, y = y)) +
geom_point(color = "#2E8B57", alpha = 0.6, size = 2) +
geom_smooth(method = "lm", se = FALSE, color = "#D55E00",
linetype = "dashed", linewidth = 0.8) +
labs(title = "Shape Scores vs Response",
subtitle = paste("R-squared =", round(fit_id$r.squared, 3)),
x = "Shape Score", y = "Response (y)")
#> `geom_smooth()` using formula = 'y ~ x'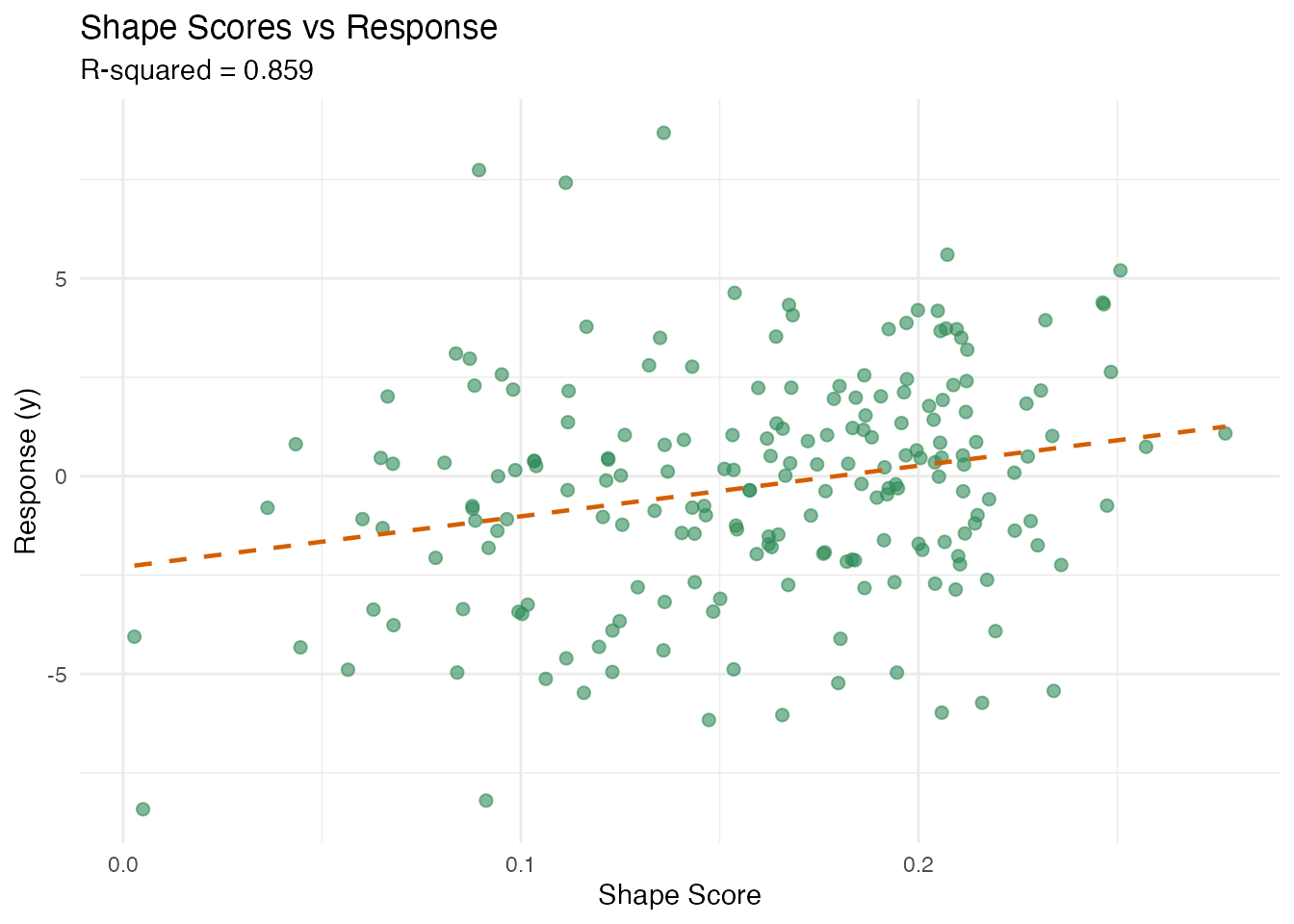
Prediction
# Hold-out evaluation
set.seed(123)
train_idx <- sample(n, 140)
test_idx <- setdiff(1:n, train_idx)
fd_train <- fd[train_idx, ]
fd_test <- fd[test_idx, ]
y_train <- y[train_idx]
y_test <- y[test_idx]
fit_train <- scalar.on.shape(fd_train, y_train, nbasis = 11, lambda = 1e-3,
index.method = "identity",
max.iter.outer = 15, tol = 1e-4)
y_pred <- predict(fit_train, fd_test)
cat("Test RMSE:", round(sqrt(mean((y_test - y_pred)^2)), 4), "\n")
#> Test RMSE: 1.035
cat("Test R2:", round(1 - sum((y_test - y_pred)^2) /
sum((y_test - mean(y_test))^2), 4), "\n")
#> Test R2: 0.8463Comparison with Other Methods
We compare three regression approaches on our phase-variable data. The key question is: can each method extract the shape signal (bump width) when the dominant variation is phase (bump location)?
Standard Regression (Ignoring Phase)
fregre.lm() treats the raw (misaligned) curves as input.
Because FPCA captures the dominant phase variation first, the
shape signal that drives
is buried in higher-order components. With a practical choice of 5
components, fregre.lm struggles to find the signal.
fit_std <- fregre.lm(fd_train, y_train, ncomp = 5)
pred_std <- predict(fit_std, fd_test)
cat("fregre.lm R-squared (train):", round(fit_std$r.squared, 4), "\n")
#> fregre.lm R-squared (train): 0.408
cat("fregre.lm RMSE (test):",
round(sqrt(mean((y_test - pred_std)^2)), 4), "\n")
#> fregre.lm RMSE (test): 97.5319To understand why fregre.lm() fails, consider
the FPCA decomposition: the first few eigenfunctions capture
where the bump is (phase), not how wide it is (shape).
The shape information only appears in higher-order components that are
typically discarded.
Elastic Regression (Alignment-Aware)
elastic.regression() jointly aligns curves and fits a
coefficient function. It captures amplitude variation after alignment
but is not specifically designed to isolate shape features.
fit_elastic <- elastic.regression(fd_train, y_train, ncomp.beta = 5,
lambda = 0.01, max.iter = 20, tol = 1e-3)
pred_elastic <- predict(fit_elastic, fd_test)
cat("elastic.regression R-squared (train):",
round(fit_elastic$r.squared, 4), "\n")
#> elastic.regression R-squared (train): 0.0418
cat("elastic.regression RMSE (test):",
round(sqrt(mean((y_test - pred_elastic)^2)), 4), "\n")
#> elastic.regression RMSE (test): 1.7091Summary Table
pred_sos <- predict(fit_train, fd_test)
rmse_std <- sqrt(mean((y_test - pred_std)^2))
rmse_elastic <- sqrt(mean((y_test - pred_elastic)^2))
rmse_sos <- sqrt(mean((y_test - pred_sos)^2))
r2_test <- function(actual, predicted) {
1 - sum((actual - predicted)^2) / sum((actual - mean(actual))^2)
}
comparison <- data.frame(
Method = c("fregre.lm (ncomp=5)",
"elastic.regression",
"scalar.on.shape"),
Handles_Phase = c("No", "Yes (alignment)", "Yes (shape score)"),
Train_R2 = round(c(fit_std$r.squared,
fit_elastic$r.squared,
fit_train$r.squared), 4),
Test_RMSE = round(c(rmse_std, rmse_elastic, rmse_sos), 4),
Test_R2 = round(c(r2_test(y_test, pred_std),
r2_test(y_test, pred_elastic),
r2_test(y_test, pred_sos)), 4)
)
knitr::kable(comparison, caption = "Method comparison on phase-variable data")| Method | Handles_Phase | Train_R2 | Test_RMSE | Test_R2 |
|---|---|---|---|---|
| fregre.lm (ncomp=5) | No | 0.4080 | 97.5319 | -1363.4010 |
| elastic.regression | Yes (alignment) | 0.0418 | 1.7091 | 0.5810 |
| scalar.on.shape | Yes (shape score) | 0.8312 | 1.0350 | 0.8463 |
scalar.on.shape() achieves the best predictive accuracy
because it directly targets the phase-invariant shape feature. Standard
regression with 5 components captures almost none of the signal, while
elastic regression provides an intermediate result.
Visualizing Predictions
df_pred <- data.frame(
Observed = rep(y_test, 3),
Predicted = c(pred_std, pred_elastic, pred_sos),
Method = factor(rep(c("fregre.lm (ncomp=5)", "elastic.regression",
"scalar.on.shape"),
each = length(y_test)),
levels = c("fregre.lm (ncomp=5)", "elastic.regression",
"scalar.on.shape"))
)
ggplot(df_pred, aes(x = Observed, y = Predicted)) +
geom_point(alpha = 0.7, color = "#4A90D9") +
geom_abline(slope = 1, intercept = 0, linetype = "dashed", color = "gray50") +
facet_wrap(~ Method) +
labs(title = "Observed vs Predicted by Method",
x = "Observed", y = "Predicted")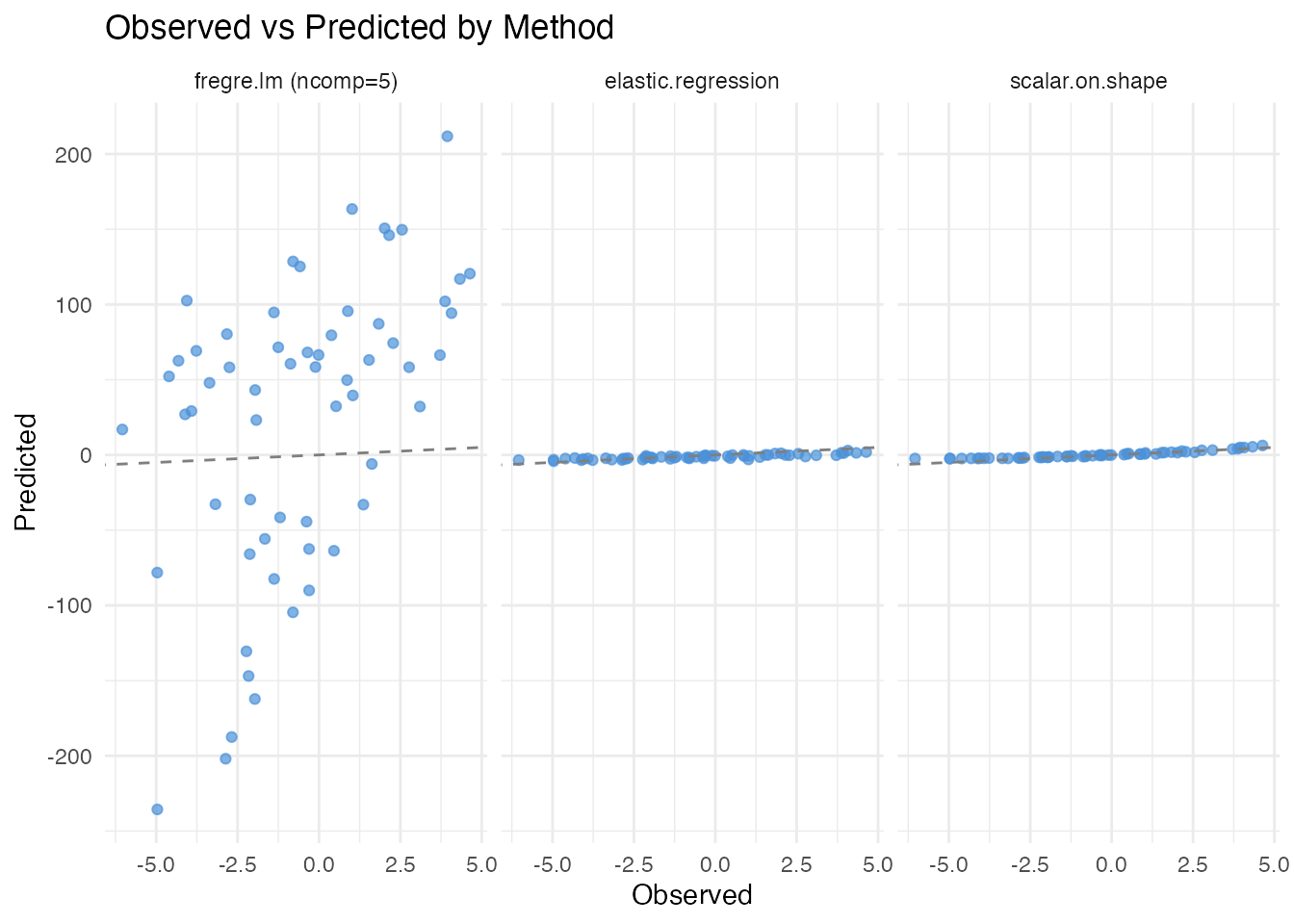
Why Standard Regression Fails
To see why more components don’t fully solve the problem for
fregre.lm(), consider what happens as we increase
ncomp:
ncomp_vals <- c(3, 5, 8, 12)
r2_sweep <- sapply(ncomp_vals, function(k) {
fit <- fregre.lm(fd_train, y_train, ncomp = k)
pred <- predict(fit, fd_test)
r2_test(y_test, pred)
})
df_sweep <- data.frame(ncomp = ncomp_vals, test_r2 = r2_sweep)
ggplot(df_sweep, aes(x = ncomp, y = test_r2)) +
geom_line(color = "#4A90D9", linewidth = 1) +
geom_point(color = "#4A90D9", size = 3) +
geom_hline(yintercept = r2_test(y_test, pred_sos),
linetype = "dashed", color = "#D55E00") +
annotate("text", x = max(ncomp_vals), y = r2_test(y_test, pred_sos),
label = "scalar.on.shape", vjust = -0.5, color = "#D55E00") +
labs(title = "fregre.lm Performance vs Number of Components",
x = "Number of FPC Components", y = expression("Test R"^2)) +
ylim(-0.2, 1)
#> Warning: Removed 4 rows containing missing values or values outside the scale range
#> (`geom_line()`).
#> Warning: Removed 4 rows containing missing values or values outside the scale range
#> (`geom_point()`).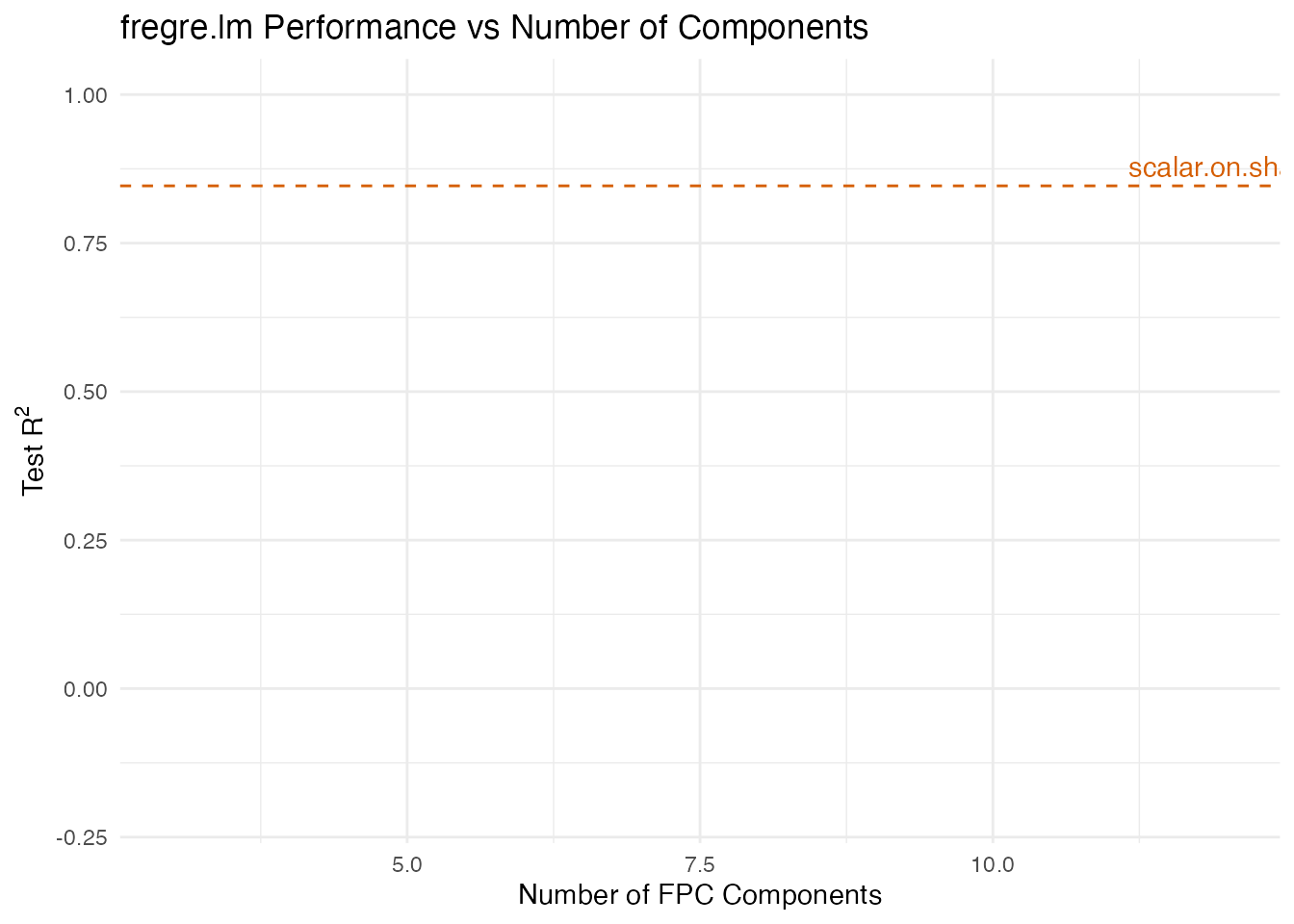
fregre.lm() eventually catches up with many components,
but requires 8–12 to begin matching scalar.on.shape(). In
practice, selecting the right number of components is difficult, and
using too many risks overfitting. The phase-invariant approach avoids
this model selection problem entirely.
Tuning Parameters
| Parameter | Default | Role | Guidance |
|---|---|---|---|
nbasis |
11 | Fourier basis dimension for | Increase for complex shapes; odd values recommended |
lambda |
1e-3 | Roughness penalty on | Increase to smooth ; decrease to capture detail |
index.method |
"identity" |
Link function type | Start with identity; try polynomial if nonlinear |
index.param |
2 | Polynomial degree or NW bandwidth | Higher = more flexible (risk of overfitting) |
max.iter.outer |
10 | Outer loop iterations | 10–20 usually sufficient |
tol |
1e-4 | Convergence threshold (relative SSE change) | Tighter = more iterations, better fit |
References
Srivastava, A., Klassen, E., Joshi, S.H. and Jermyn, I.H. (2011). Shape Analysis of Elastic Curves in Euclidean Spaces. IEEE Transactions on Pattern Analysis and Machine Intelligence, 33(7), 1415–1428.
Tucker, J.D., Wu, W. and Srivastava, A. (2013). Generative Models for Functional Data Using Phase and Amplitude Separation. Computational Statistics & Data Analysis, 61, 50–66.
See Also
-
vignette("articles/elastic-regression")— elastic scalar-on-function regression and logistic classification -
vignette("articles/scalar-on-function")— standard scalar-on-function regression methods -
vignette("articles/elastic-alignment")— curve alignment without regression